
Genetics: Analysis and Principles
6th Edition
ISBN: 9781259616020
Author: Robert J. Brooker Professor Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 15, Problem 6EQ
As described in Chapter 21, an electrophoretic mobility shift assay (EMSA) can be used to determine if a protein binds to a segment of DNA. When a segment of DNA is bound by a protein, its mobility will be retarded, and the DNA band will appear higher in the gel. In an EMSA whose results are shown below, a cloned gene fragment that is
in length contains a regulatory element that is recognized by a transcription factor called protein X. Previous experiments have shown that the presence of hormone X results in transcriptional activation by protein X.
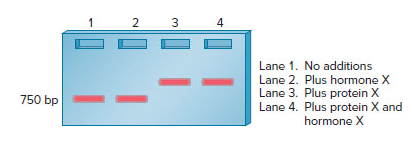
Explain the action of hormone X.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
GR and PPAR are transcription factors that bind to GRE and PPARE sequences respectively and activate transcription of genes. A reporter cell line is created in which the the green fluoresecent protein (GFP) is controlled by a GRE sequence and the pink fluorescent protein mCherry is under control of a PPARE sequence. If the gene for GR is introduced into the reporter cell line, the cells produce a green color. Chimeric proteins are created in which the DNA Binding Domains (DBD) and Activation Domains (AD) of the transcription factors are introduced into various cell lines. Match the following cell-types with the fluorescent color(s) you would expect the cells to produce.
In cells, certain hormones, such as epinephrine, have the ability to raise the concentration of cAMP. The CREB protein can be found in a cell extract that has been pretreated with epinephrine or not, depending on your preference.Using an electrophoretic mobility shift experiment, you test the CREB protein's capacity to attach to a DNA fragment containing a cAMP response element (CRE). Specify what you hope to achieve.
(c) By binding one L-tryptophan molecule/monomer, the trp repressor binds to DNA to suppress syn-
thesis of L-tryptophan in E. coli. Below is the amino acid sequence of the helix – (reverse) turn – helix
region of the trp repressor that binds to DNA compared to the sequence of the corresponding DNA
binding motif of the Prl protein, a different type of repressor protein. A diagram of the trp repressor
dimer is also shown.
reverse turn
trp helix 4
70
Trp
-Gly-Glu-Met-Ser-Gln-Arg-Glu-Leu-Lys-Asn-Glu-Leu-Gly-Ala-Gly-
Ile-
Prl
-Ser-Glu-Glu-Ala-Lys-Glu-Glu-Leu-Ala-Lys-Lys-Cys-Gly-Ile-Thr-
Val-
Pri heilix
trp helix 5
80
90
Trp
Ala-Thr-Ile-Thr-Arg-Gly-Ser sgn-Ser-Leu-Lys-Ala-Ala-
Prl
Ser-Gln-Val-Ser-Asn-Trp-Phe-Gly-Asn-Lys-Arg-Ile-Arg-
Prl helix
Chapter 15 Solutions
Genetics: Analysis and Principles
Ch. 15.1 - 1. Combinatorial control refers to the phenomenon...Ch. 15.1 - 2. A regulatory transcription factor protein...Ch. 15.1 - 3. A bidirectional enhancer has the following...Ch. 15.1 - 4. Regulatory transcription factors can be...Ch. 15.2 - 1. A chromatin-remodeling complex may
a. change...Ch. 15.2 - Prob. 2COMQCh. 15.2 - 3. Which of the following characteristics is...Ch. 15.2 - 4. Transcriptional activation of eukaryotic genes...Ch. 15.3 - How can methylation affect transcription? a. It...Ch. 15.3 - 2. The process in which completely unmethylated...
Ch. 15.4 - Prob. 1COMQCh. 15.5 - The overall goal of the ENCODE Project is a. to...Ch. 15.6 - The binding of iron regulatory protein (IRP) to...Ch. 15 - Discuss the common points of control in eukaryotic...Ch. 15 - 2. Discuss the structure and function of...Ch. 15 - 3. What is meant by the term transcription factor...Ch. 15 - What are the functions of transcriptional...Ch. 15 - 5. Is each of the following statements true or...Ch. 15 - 6. Transcription factors usually contain one or...Ch. 15 - Prob. 7CONQCh. 15 - Prob. 8CONQCh. 15 - 9. Let’s suppose a mutation in the glucocorticoid...Ch. 15 - Prob. 10CONQCh. 15 - Prob. 11CONQCh. 15 - Prob. 12CONQCh. 15 - 13. Transcription factors such as the...Ch. 15 - An enhancer, located upstream from a gene, has the...Ch. 15 - 15. The DNA-binding domain of each CREB protein...Ch. 15 - The gene that encodes the enzyme called tyrosine...Ch. 15 - Prob. 17CONQCh. 15 - 18. What is a histone variant?
Ch. 15 - Prob. 19CONQCh. 15 - 20. What is meant by the term histone code? With...Ch. 15 - Prob. 21CONQCh. 15 - Histones are thought to be displaced as RNA...Ch. 15 - 23. What is an insulator? Describe two different...Ch. 15 - 24. What is DNA methylation? When we say that DNA...Ch. 15 - Lets suppose that a vertebrate organism carries a...Ch. 15 - 26. What is a CpG island? Where would you expect...Ch. 15 - Describe how the binding of iron regulatory...Ch. 15 - 1. Briefly describe the method of chromatin...Ch. 15 - Researchers can isolate a sample of cells, such as...Ch. 15 - Prob. 3EQCh. 15 - Prob. 4EQCh. 15 - Prob. 5EQCh. 15 - 6. As described in Chapter 21, an electrophoretic...Ch. 15 - Prob. 7EQCh. 15 - 1. Explain how DNA methylation could be used to...Ch. 15 - 2. Enhancers can occur almost anywhere in DNA and...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- The interphase nucleus is a highly structured organelle with chromosome territories, interchromatin compartments, and transcription factories. In cultured human cells, researchers have identified approximately 8000 transcription factories per cell, each containing an average of eight tightly associated RNAP II molecules actively transcribing RNA. If each RNAP II molecule is transcribing a different gene, how might such a transcription factory appear? Provide a simple diagram that shows eight different genes being transcribed in a transcription factory and include the promoters, structural genes, and nascent transcripts in your presentation.arrow_forwardNegative supercoiling of DNA favors the transcription of genes because it facilitates unwinding. However, not all promoter sites are stimulated by negative supercoiling. The promoter site for topoisomerase II itself is a noteworthy exception. Negative supercoiling decreases the rate of transcription of this gene. Propose a possible mechanism for this effect and suggest a reason why it may occur.arrow_forwardc) You use chromatin immunoprecipitation to measure the location and amount of transcription factors binding around the Lrg1 gene in the four treatments. You use your data to generate the following figure. Orient yourself to the figure by finding the transcription start site on the x-axis. The size of the peak indicates the frequency of the transcription factor at that position. Lrg1 Lrg1 transcription Control u = 12 2. 3 Sucrose = 77 Sucrose + = 180 Hormone 4 Starvation = 2 -2,000 bp -1,000 bp +1 bp +1,000 bp i) What type of transcription factor might be present at position 2? What evidence from the figure supports your claim? (two sentences max) What type of transcription factor might be present at position 3? What evidence ii) from the figure supports your claim? (two sentences max) iii) Look at the transcription factors at position 4. Use evidence from the figure to explain the effect they are having on transcription of Lrg1. (two sentences max)arrow_forward
- 3′-->5′ Exonuclease activity allows DNA polymerase III (Pol III) to back-up and fix a mismatched base pair that was just incorporated into a growing strand of new DNA. True Or False In one of the four ways to regulate gene expression, positive control with repression indicates that transcription is activated in the presence of a co-repressor. True Or Falsearrow_forwardThe chart below is a position specific scoring matrix (PSSM, a logarithmic transformed matrix) for a transcription factor binding site. (1). Evaluate a sequence “GACATTCA” to find out which segment of the sequence fits the binding site best. (2) What is the max score that a sequence can have with this PSSM? (3) What is the minimum score a sequence can have with this PSSM?arrow_forwardTo identify the following types of genetic occurrences, would acomputer program use sequence recognition, pattern recognition,or both?A. Whether a segment of Drosophila DNA contains a P element(which is a specific type of transposable element)B. Whether a segment of DNA contains a stop codonC. In a comparison of two DNA segments, whether there is aninversion in one segment compared with the other segmentD. Whether a long segment of bacterial DNA contains one ormore genesarrow_forward
- You would like to add a nuclear localization sequence (NLS) of Lys-Lys-Lys-Arg-Lys to a protein that is usually found in the cytoplasm of a yeast cell. To accomplish this, you introduce the nucleotide sequence encoding the NLS into the gene that encodes the cytoplasmic protein of interest. a. What is the size of the nucleotide insert that will encode the NLS? Briefly explain. 5' 3' b. Below is a diagram of the gene encoding the cytoplasmic protein of interest in the yeast genome. If your goal is to put the NLS at the carboxyl (C) terminus of the protein, at which location (A-E) should the NLS be inserted? Briefly explain. A TATAA ATATT promoter +1 B ATG TAC D TAA ATT stop codon E 3' 5'arrow_forwardThe following diagram show what is required for an active promoter of a gene of interest, where:A1 = Activator 1A2 = Activator 2Med = MediatorRep = Repressor Based on the following data, predict: Chromatin conformation Methylation state of the proximal promoter If protein A1 is present or absent If protein A2 is present or absent If the mediator is present or absent If the repressor is absence or present If this gene is likely to be transcribed or notPlease note that this is an "all or nothing" bonus question, and no partial credit will be awarded. Selecting all answers will result in zero points awarded. Selecting at least one incorrect answer will result in zero points being awarded. Question 28 options: Euchromatin Heterochromatin Methylated promoter Unmethylated promoter Activator A1 present Activator A1 absent Activator A2 present…arrow_forwardWhen Laybourne and Kadonaga studied the effects of histone proteins on eukaryotic transcription using an in vitro transcription assay explain why: a) they used two different DNA templates that contained different promoter structures. b) when they included both activator protein and histones, they always added the histone proteins before adding the activator to the transcription assay mixture. (Ctri) -arrow_forward
- For a specific type of mutation at a given location in a particular gene, identify whether it will affect the size of the mRNA, the protein, or both. How would the mutant appear on a gel in comparison to the originalarrow_forwardChromatin Immunoprecipitation (ChIP) experiments enable researchers to measure the levels of transcription factors, coactivators/corepressors and chromatin remodeling complexes on a specific gene in cells. In the ChIP experiment below, the recruitment of the transcription factor NfkB, Histone Deacetylase Complex (HDAC3) and p300/CBP complex to the Interleukin 12 (IL12) gene at various times (0, 30, 60 and 120 min) after treating cells with LPS are measured. Does LPS stimulate or inhibit transcription of the IL12 gene based on the recruitment of factors to the IL12 gene. Justify your answer by explaining the state of the gene before LPS and after LPS treatment (limit 5-6 sentences).arrow_forwardThe IMD2 promoter contains three upstream transcription start sites (TSS) that are utilized under high GTP conditions and a single downstream TSS (-106) that is normally only utilized under low GTP conditions. In a wild type cell, expression of IMD2 mRNA only occurs if transcription initiates from the -106 TSS. In 300 words or less, describe: 1.) The normal function of Ssl2, and 2.) why a mutation in Ssl2, that increases its catalytic rate, would allow expression of the IMD2 ORF under high GTP conditions. (Conditions under which the IMD2 ORF is NOT expressed in the wild type.)arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
QCE Biology: Introduction to Gene Expression; Author: Atomi;https://www.youtube.com/watch?v=a7hydUtCIJk;License: Standard YouTube License, CC-BY