
Concepts of Genetics (12th Edition)
12th Edition
ISBN: 9780134604718
Author: William S. Klug, Michael R. Cummings, Charlotte A. Spencer, Michael A. Palladino, Darrell Killian
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 20, Problem 29ESP
The gel presented here shows the pattern of bands of fragments produced with several restriction enzymes. The enzymes used are identified above the lanes of the gel, and six possible restriction maps are shown in the column to the right.
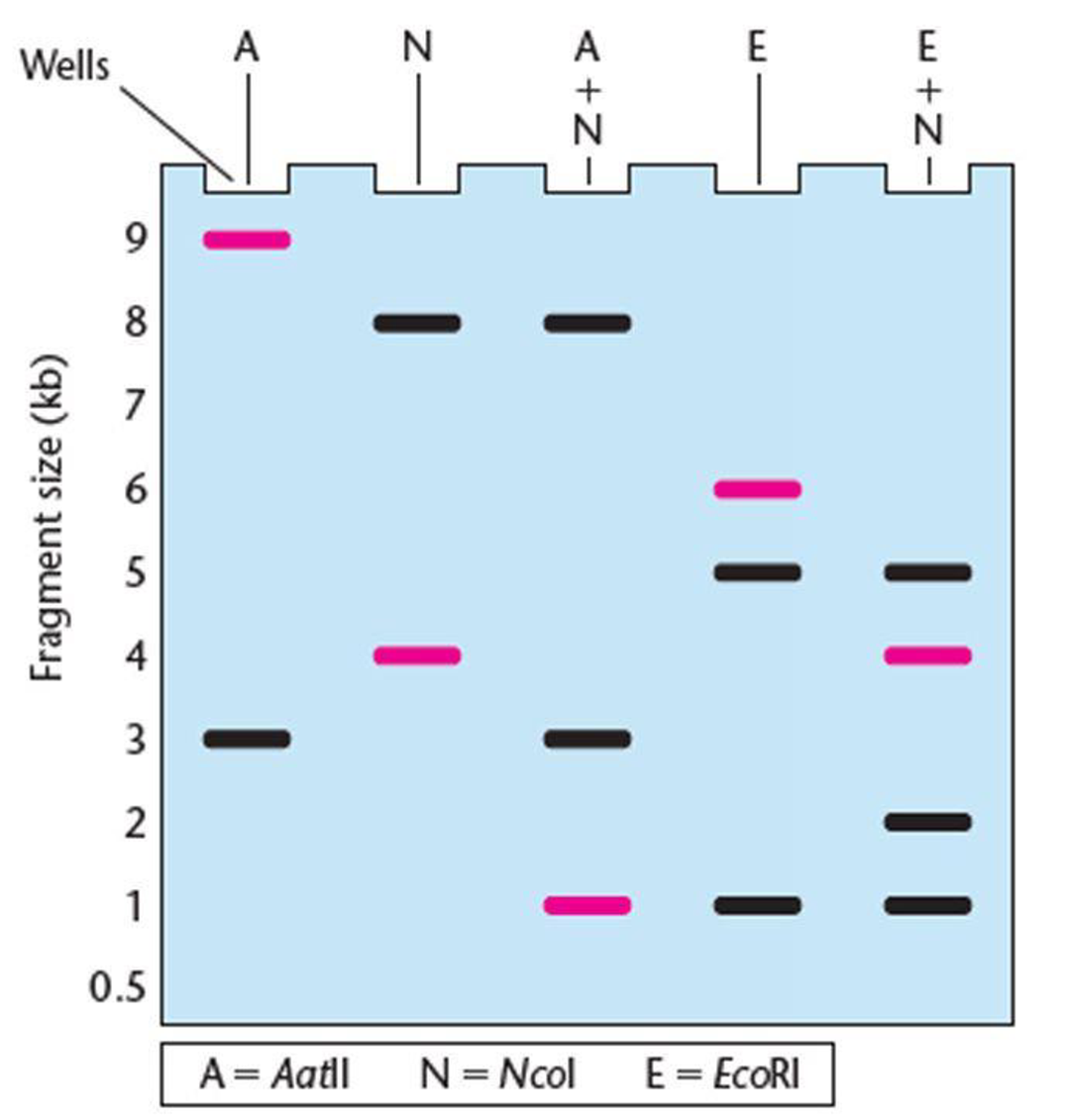
One of the six restriction maps shown is consistent with the pattern of bands shown in the gel.
- (a) From your analysis of the pattern of bands on the gel, select the correct restriction map and explain your reasoning.
- (b) The highlighted bands (magenta) in the gel hybridized with a probe for the gene pep during a Southern blot. Where in the gel is the pep gene located?
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
A small DNA molecule was cleaved with several different restriction nucleases, and the size of each fragment was determined by gel electrophoresis.The following data were obtained.
(a) Is the original molecule linear or circular?(b) Draw a map of restriction sites (showing distances between sites) that isconsistent with the data given.(c) How many additional maps are compatible with the data?(d) What would have to be done to locate the cleavage sites unambiguouslywith respect to each other?
If a 1000 bp of DNA were inserted between the two restriction sites, how would the banding pattern on the gel differ from the one you drew in part a?
(PART A WITH THE FIRST PART OF THE QUESTION IS ATTACHED)
Table 21.3 describes the cleavage sites of five different restrictionenzymes. After these restriction enzymes have cleaved the DNA, four of them produce sticky ends that can hydrogen bond with complementary sticky ends, as shown in Figure 21.1. The efficiency of sticky ends binding together depends on the number of hydrogen bonds; more hydrogen bonds makes the ends “stickier” and more likely to stay attached. Rank these four restriction enzymes from Table 21.3 (from best to worst)with regard to the efficiency of their sticky ends binding to each other.
Chapter 20 Solutions
Concepts of Genetics (12th Edition)
Ch. 20 - A plasmid that is both ampicillin and tetracycline...Ch. 20 - You have just created the worlds first genomic...Ch. 20 - What undesirable or unforeseen consequences might...Ch. 20 - Do we have the ethical right to alter the genomes...Ch. 20 - Should these new technologies be regulated...Ch. 20 - HOW DO WE KNOW? In this chapter we focused on how...Ch. 20 - CONCEPT QUESTION Review the Chapter Concepts list...Ch. 20 - What roles do restriction enzymes, vectors, and...Ch. 20 - The human insulin gene contains a number of...Ch. 20 - Although many cloning applications involve...
Ch. 20 - Using DNA sequencing on a cloned DNA segment, you...Ch. 20 - Restriction sites are palindromic; that is, they...Ch. 20 - List the advantages and disadvantages of using...Ch. 20 - What are the advantages of using a restriction...Ch. 20 - In 1975, the Asilomar Conference on Recombinant...Ch. 20 - In the context of recombinant DNA technology, of...Ch. 20 - If you performed a PCR experiment starting with...Ch. 20 - Prob. 13PDQCh. 20 - Prob. 14PDQCh. 20 - You have recovered a cloned DNA segment from a...Ch. 20 - Prob. 16PDQCh. 20 - Although the capture and trading of great apes has...Ch. 20 - Prob. 18PDQCh. 20 - Prob. 19PDQCh. 20 - Prob. 20PDQCh. 20 - Traditional Sanger sequencing has largely been...Ch. 20 - How is fluorescent in situ hybridization (FISH)...Ch. 20 - What is the difference between a knockout animal...Ch. 20 - Prob. 24PDQCh. 20 - When disrupting a mouse gene by knockout, why is...Ch. 20 - Prob. 26PDQCh. 20 - Prob. 27PDQCh. 20 - As you will learn later in the text (Special...Ch. 20 - The gel presented here shows the pattern of bands...Ch. 20 - A widely used method for calculating the annealing...Ch. 20 - Most of the techniques described in this chapter...Ch. 20 - In humans, congenital heart disease is a common...Ch. 20 - The U.S. Department of Justice has established a...Ch. 20 - Prob. 34ESP
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- A 12 kb linear DNA fragment is subject to single or double RE digest and agarose gelelectrophoresis, to yield the gel profile shown below. The first lane contains the size marker(M).a) Explain how the name of the enzyme EcoRI is derived.b) How many sites are there for EcoRI and PvuII respectively on this DNA fragment?c) Use the sizes of the DNA bands on the gel to compile a restriction enzyme map of the DNAfragment. Indicate the positions of the restriction enzymes sites for EcoRI and PvuII on themap.arrow_forwardGiven the DNA sequence of the restriction enzyme: gi|6329444|dbj|AB034757.1| Hynobius retardatus mRNA for larval beta-globin, complete cds GCAGAATCTGACTCAAGAAATCCCTCCTCACCCAACACCACCAGCAGCCATGGTTCACTGGACAGCAGAGGAGAAGGCAGCCATCAGCTCTGTGTGGAAGCAGGTGAACGTGGAGAGCGATGGACAGGAGGCCCTGGCCAGGTTGCTGATCGTCTACCCCTGGACCCAGAGATACTTCAGCTCTTTTGGGGACCTGTCGAGCCCAGCTGCCATTTGTGCCAACGCCAAGGTCCGTGCCCATGGCAAGAAGGTCCTGTCCGCCCTGGGAGCCGGCGCCAACCACCTGGATGACATCAAAGGCAACTTTGCTGATCTGAGCAAGCTTCACGCAGACACACTCCATGTGGACCCCAATAACTTCCTGCTCCTGGCAAACTGCCTGGTGATCGTCTTGGCCCGCAAGCTGGGAGCCGCCTTCAACCCTCAAGTCCATGCGGCCTGGGAGAAGTTCCTGGCCGTCTCCACCGCGGCTCTGTCCAGAAACTACCACTAGAGACTGGTCTTTGGGTTTAATTCTGTGAACGTCCCTGAGACAAATGATCTTTCAATGTGTAAACCTGTCATTACATCAATAAAGAGACATCTAACAAAAAAAAAAAAAAAAAAAAAAAAAA Identify two blunt-end cutters Identify two sticky-end cutters. For each, Provide the sequence of the Restriction enzyme, Highlight using a specific color where the DNA sequence where the restriction enzyme will cut the DNA Indicate the…arrow_forwardIf a 500 bp of DNA between the two restriction sites were deleted, how would the banding pattern on the gel differ from the one you drew in part a? (Part a is attached)arrow_forward
- 1) Restriction enzymes come in a concentration of U/ml, and it is recommended that 1U be used for each ug of DNA to be digested. If an enzyme you want to use comes in at 22,000 U/ml, and you want to digest 5 ug of DNA. How much volume will you have to use for the reaction and how will you be able to measure it with the pipettes we have in the lab? 2) An enzyme for ligation (Cip) comes at a concentration of 16000 units/ml. How many units will there be in 10 ul?arrow_forwardUsing the data in Table, identify restriction enzymes that (a) produce blunt ends; (b) recognize and cleave the same sequence (called isoschizomers); (c) produce identical sticky ends.arrow_forwardA plasmid DNA and a linear DNA (both of the same size) have one site for a restriction endonuclease. When cut and separated on agarose gel electrophoresis, plasmid shows one DNA band while linear DNA shows two fragments. Explain.arrow_forward
- The sequences below indicated the 6bp recognition site for the restriction enzyme EcoRI. The lines indicate the sites where the enzyme will cut each strand. 1). write the sequence and structure of the two DNA pieces after the enzyme cuts (hydrogen bonds holding the strands together between the lines are broken after enzyme cuts) 2). indicate whether EcoRI generates blunt or sticky overhangs 5'- G I A A T T C - 3' 3' - C T T A A l G - 5'arrow_forwardYou are studying a new plasmid, and you digest the plasmid with three restriction enzymes: Eco RI (E), HindlII (H), and Xbal (X). You digest the plasmid DNA with each of the following combinations of enzymes and observe the results on an agarose gel. You are provided a partial plasmid map as shown below to the right. E H E+H E+X H+X Kb +4.3 +2.8 +2.5 - +2.0 -- -1.8 -1.5 12 +1.0 +0.8 +0.5 - a. What is the size of this plasmid in base pairs? b. What is the distance in base pairs between E1 and H? c. What is the distance in base pairs between E1 and X? d. What is the distance in base pairs between E2 and H? e. What is the distance in base pairs between E2 and X?arrow_forwardOn the gel shown below are four DNA samples. Samples A to C are taken from tissues of landslide victims that are being identified, while sample D came from a hair sample brought by a mother looking for the remains of her son. (see img) i. If similar band patterns in a gel are created using the same restriction enzyme, what does that tell you about the DNA sequence of the samples? ii. In sample C, only two fragments were created. How many restriction sites (regions where enzymes cut) are present in sample C?arrow_forward
- A group of overlapping clones, designated A through F, is isolated from one region of a chromosome. Each of the clones is separately cleaved by a restriction enzyme, and the pieces are resolved by agarose gel lectrophoresis,with the results shown below. There are nine different restriction fragments in this chromosomal region, with a subset appearing in each clone. Using this information, deduce the order of the restriction fragments in the chromosome.arrow_forwardAfter restriction enzymes cut, they contain unpaired bases. Type II restriction enzymes leave ends that may be 5' overhanging, 3' overhanging, or blunt. In all cases each end is left with a 3' OH and a 5' phosphate. All blunt ends, and any complementary overhanging ends may be re-ligated with T4 DNA ligase, as long as at least one 5'- phosphate is present. In the tables below G^AATTC means that the end after cutting with enzyme will be: -----G 3' -----CTTAA 5' GTGCA^C means that the end will be: -----GTGCA 3' -----C 5' Which RE’s from table below have a 5’ overhang? Which ones have a 3’ Overhang? AccI GT^CGAC BamHI G^GATCC ClaI AT^CGAT NsiI ATGCA^T PstI CTGCA^G BglII A^GATCT TaqI T^CGAarrow_forwardYou are studying a new plasmid, and you digest the plasmid with three restriction enzymes: Eco RI (E), HindlII (H), and Xbal (X). You digest the plasmid DNA with each of the following combinations of enzymes and observe the results on an agarose gel. You are provided a partial plasmid map as shown below to the right. E+H E+X H+x Kb +4.3 +2.8 -+2.5 -2.0 - -1.8 +1.5 -1.0 12 F0.8 +0.5 a. What is the size of this plasmid in base pairs? b. What is the distance in base pairs between E1 and H? c. What is the distance in base pairs between E1 and X? d. What is the distance in base pairs between E2 and H? e. What is the distance in base pairs between E2 and X?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Bacterial Genomics and Metagenomics; Author: Quadram Institute;https://www.youtube.com/watch?v=_6IdVTAFXoU;License: Standard youtube license