
Concepts of Genetics (12th Edition)
12th Edition
ISBN: 9780134604718
Author: William S. Klug, Michael R. Cummings, Charlotte A. Spencer, Michael A. Palladino, Darrell Killian
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 20, Problem 15PDQ
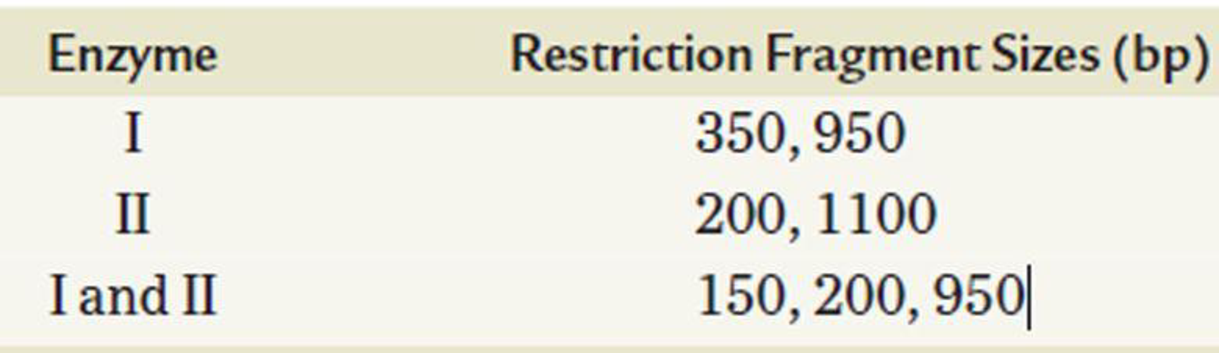
You have recovered a cloned DNA segment from a vector and determine that the insert is 1300 bp in length. To characterize this cloned segment, you isolate the insert and decide to construct a restriction map. Using enzyme I and enzyme II, followed by gel electrophoresis, you determine the number and size of the fragments produced by enzymes I and II alone and in combination, as recorded in the following table. Construct a restriction map from these data, showing the positions of the restriction-enzyme cutting sites relative to one another and the distance between them in units of base pairs.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Below is a diagram of the vector you are
planning to use. You identify four restriction
enzyme recognition sites in the vector as
indicated in the diagram below. The distance
between the restriction sites are indicated
by numbers (in kilo base pairs).
ori
ВатHI
ВатHI
2
ЕcoRI
Kpnl
If you digest the vector with a combination
of EcoRI and BamHI and run the resulting
DNA fragments on a gel, which lane would
represent the expected result? (Lane 1
contains the DNA size marker.)
А В с
(kbp)
8
Promoter
A small DNA molecule was cleaved with several different restriction nucleases, and the size of each fragment was determined by gel electrophoresis. The following data were obtained.
Single and double digestion of plasmid pMCS326 were performed using the restriction enzymes AluIII and EcoRV. DNA fragments were shown in an electrophoretogram below. Construct a restriction map of plasmid pMCS326 for enzymes AluIII and EcoRV. (Create restriction mapping with explanation)
Chapter 20 Solutions
Concepts of Genetics (12th Edition)
Ch. 20 - A plasmid that is both ampicillin and tetracycline...Ch. 20 - You have just created the worlds first genomic...Ch. 20 - What undesirable or unforeseen consequences might...Ch. 20 - Do we have the ethical right to alter the genomes...Ch. 20 - Should these new technologies be regulated...Ch. 20 - HOW DO WE KNOW? In this chapter we focused on how...Ch. 20 - CONCEPT QUESTION Review the Chapter Concepts list...Ch. 20 - What roles do restriction enzymes, vectors, and...Ch. 20 - The human insulin gene contains a number of...Ch. 20 - Although many cloning applications involve...
Ch. 20 - Using DNA sequencing on a cloned DNA segment, you...Ch. 20 - Restriction sites are palindromic; that is, they...Ch. 20 - List the advantages and disadvantages of using...Ch. 20 - What are the advantages of using a restriction...Ch. 20 - In 1975, the Asilomar Conference on Recombinant...Ch. 20 - In the context of recombinant DNA technology, of...Ch. 20 - If you performed a PCR experiment starting with...Ch. 20 - Prob. 13PDQCh. 20 - Prob. 14PDQCh. 20 - You have recovered a cloned DNA segment from a...Ch. 20 - Prob. 16PDQCh. 20 - Although the capture and trading of great apes has...Ch. 20 - Prob. 18PDQCh. 20 - Prob. 19PDQCh. 20 - Prob. 20PDQCh. 20 - Traditional Sanger sequencing has largely been...Ch. 20 - How is fluorescent in situ hybridization (FISH)...Ch. 20 - What is the difference between a knockout animal...Ch. 20 - Prob. 24PDQCh. 20 - When disrupting a mouse gene by knockout, why is...Ch. 20 - Prob. 26PDQCh. 20 - Prob. 27PDQCh. 20 - As you will learn later in the text (Special...Ch. 20 - The gel presented here shows the pattern of bands...Ch. 20 - A widely used method for calculating the annealing...Ch. 20 - Most of the techniques described in this chapter...Ch. 20 - In humans, congenital heart disease is a common...Ch. 20 - The U.S. Department of Justice has established a...Ch. 20 - Prob. 34ESP
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- A piece of DNA 5.0 kb long is cloned and then cut out of the vector for analysis. This linear piece of DNA is digested with two restriction enzymes, EcoRI and BamHI, individually and in combination, and the resulting fragment sizes are determined by electrophoresis. The results are as follows: Restriction fragment size 4.5 kb; 0.5 kb 3.0 kb; 2.0 kb 2.5 kb; 2.0 kb; 0.5 kb Enzyme name EcoRI ВатHI EcoRI + BamHI Construct a potential restriction map based on these results.arrow_forwardConstruct a detail linear restriction map according to the given information in Table 1 below. Table 1 Restriction enzyme EcoRI No, of fragments 2 Size of fragment (kb) 2,9 ВатHI 3 6, 1,4 Xhol 8, 3 EcoRI + BamHI 4 2, 4, 1,4 EcoRI + Xhol 3 2, 6, 3arrow_forwardYou are studying a new plasmid, and you digest the plasmid with three restriction enzymes: Eco RI (E), HindlII (H), and Xbal (X). You digest the plasmid DNA with each of the following combinations of enzymes and observe the results on an agarose gel. You are provided a partial plasmid map as shown below to the right. E H E+H E+X H+X Kb +4.3 +2.8 +2.5 - +2.0 -- -1.8 -1.5 12 +1.0 +0.8 +0.5 - a. What is the size of this plasmid in base pairs? b. What is the distance in base pairs between E1 and H? c. What is the distance in base pairs between E1 and X? d. What is the distance in base pairs between E2 and H? e. What is the distance in base pairs between E2 and X?arrow_forward
- You are studying a new plasmid, and you digest the plasmid with three restriction enzymes: Eco RI (E), HindlII (H), and Xbal (X). You digest the plasmid DNA with each of the following combinations of enzymes and observe the results on an agarose gel. You are provided a partial plasmid map as shown below to the right. H. E+H E+X H+X Kb +4.3 -2.8 -2.5 - - -2.0 -+1.8 - +1.5 +1.0 F0.8 12 a. What is the size of this plasmid in base pairs? b. What is the distance in base pairs between E1 and H? c. What is the distance in base pairs between E1 and X? d. What is the distance in base pairs between E2 and H? e. What is the distance in base pairs between E2 and X?arrow_forwardA small DNA molecule was cleaved with several different restriction nucleases, and the size of each fragment was determined by gel electrophoresis.The following data were obtained. (a) Is the original molecule linear or circular?(b) Draw a map of restriction sites (showing distances between sites) that isconsistent with the data given.(c) How many additional maps are compatible with the data?(d) What would have to be done to locate the cleavage sites unambiguouslywith respect to each other?arrow_forwardThey have purified a 1100 bp HindIII restriction fragment that they plan to sequence. As a first step, they decided to construct a restriction map of the fragment for the enzymes EcoRI and SmaI. Below is shown an agarose gel of the appropriate digests. Draw a restriction map of the fragment and show the distances, in base pairs, between the HindIII, EcoRI, and SmaI sites. *arrow_forward
- In relation to the use of restriction enzymes in recombinant DNA technology, answer the following: You have accidentally torn the labels off two tubes (tube A and tube B), each containing a different plasmid, now you do not know which plasmid is in which tube. Fortunately, you have restriction maps for both plasmids, shown in Figure below. You have the opportunity to test just one sample from one of your tubes. By utilizing agarose gel electrophoresis technique, which restriction enzyme OR combination of restriction enzymes would you use in this experiment to determine which plasmid is found in which tube?. (Hint: if you use Hind III restriction enzyme you are going to get ONE single fragment with a molecular size of → 0.5+0.3+0.2+0.4+1+1 = 3.4 kb).arrow_forwardA group of overlapping clones, designated A through F, is isolated from one region of a chromosome. Each of the clones is separately cleaved by a restriction enzyme, and the pieces are resolved by agarose gel lectrophoresis,with the results shown below. There are nine different restriction fragments in this chromosomal region, with a subset appearing in each clone. Using this information, deduce the order of the restriction fragments in the chromosome.arrow_forwardAssume that a plasmid is 4700 base pairs in length and has restriction sites for a given restriction enzyme at the following locations: 800, 1400, 2900, and 3600. List the fragments by size that are ! expected when the plasmid is fully digested the restriction enzyme.arrow_forward
- When the restriction endonuclease EcoRI is used to digest a 10 kb DNA fragment, it produces 4 kb and 6 kb-sized fragments. Digesting the 10 kb fragment with BamHI yields three fragments, each ranging in size from one to three and a half kilobytes. Four pieces of 0.5, 1, 3 and 5.5 kb are formed after using both enzymes. Create a restriction map for this 10 kb piece of DNA using the information you have collected. Make a note of where the two enzymes cut, as well as the distances between the enzymes.arrow_forwardDNA can be cut at specific sequence motifs using enzymes called restriction endonucleases. You obtain a single PCR product from R. sphaeroides DNA that can be cut in the places shown below by solid arrows. These are the Clal specific restriction motifs. Positions of these restriction sites are indicated by nucleotide numbers in a 1023 nucleotide DNA fragment. Draw the expected band(s) for each of the bacterial species on the gel picture below if treated with the restriction enzymes as illustrated. Rhodobacter sphaeroides 2. 4. 1 0. 1023 150 627 Rhodopseudomonas palustris BisB5 0. 1023 627 Dinoroseobacter shibae DFL 12 1023 150 Silicibacter sp. TM 1040 1023 DNA ladder R sphaeroides 2. 4. 1 R palustris BisB5 D. shibae DFL12 Silicibacter sp. TM1040 1100 1000 900 800 700 600 500 400 300 200 100arrow_forwardA 12 kb linear DNA fragment is subject to single or double RE digest and agarose gelelectrophoresis, to yield the gel profile shown below. The first lane contains the size marker(M).a) Explain how the name of the enzyme EcoRI is derived.b) How many sites are there for EcoRI and PvuII respectively on this DNA fragment?c) Use the sizes of the DNA bands on the gel to compile a restriction enzyme map of the DNAfragment. Indicate the positions of the restriction enzymes sites for EcoRI and PvuII on themap.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Molecular Techniques: Basic Concepts; Author: Dr. A's Clinical Lab Videos;https://www.youtube.com/watch?v=7HFHZy8h6z0;License: Standard Youtube License