
wo techniques commonly used to study the expression patterns of genes that play a role in development are Northern blotting and in situ hybridization. As described in Chapter 21, Northern blotting is used to detect RNA that is transcribed from a particular gene. In this method, a specific RNA is detected by using a short segment of cloned DNA as a probe. The DNA probe, which is labeled, is complementary to the RNA that the researcher wishes to detect. After the DNA probe binds to the RNA within a blot of a gel, the RNA is visualized as a labeled band on a nylon membrane. For example, a DNA probe that is complementary to the bicoid mRNA could be used to specifically detect the amount of that mRNA in a blot.
A second technique, termed fluorescence in situ hybridization (FISH), is used to identify the locations of genes on chromosomes. This technique is also used to locate gene products within oocytes, embryos, and larvae. Thus, it has been commonly used by developmental geneticists to understand the expression patterns of genes during development. The micrograph in Figure 26.8b is derived from the application of the FISH technique. In this case, the probe was complementary to bicoid mRNA.
Now here is the question. Suppose a researcher has three different Drosophila strains that have loss-of-function mutations in the bicoid gene. We will call them bicoid-A, bicoid-B, and bicoid-C; the wild type is designated bicoid+. To study these mutations,
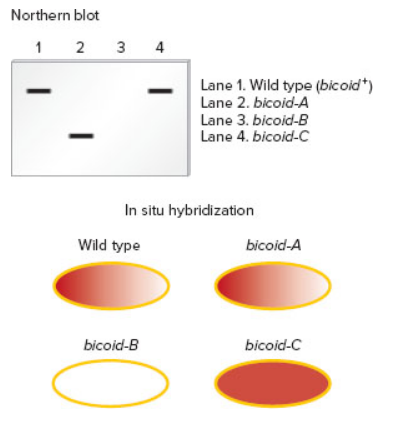
A. How can phenotypically normal female flies be homozygous for a loss-of-function allele in the bicoid gene?
B. Explain the type of mutation (e.g., deletion, point mutation, etc.) in each of the three strains. Explain how the mutation may cause a loss of normal function for the bicoid gene product.
C. Discuss how the use of both techniques provides more definitive information than the application of just one of the techniques.
Want to see the full answer?
Check out a sample textbook solution
Chapter 26 Solutions
Genetics: Analysis and Principles
- The pre-mRNA transcript and protein made by several mutant genes were examined. The results are given below. Determine where in the gene a likely mutation lies: the promoter region, exon, intron, cap on mRNA, or ribosome binding site. a. normal-length transcript, normal-length nonfunctional protein b. normal-length transcript, no protein made c. normal-length transcript, normal-length mRNA, short nonfunctional protein d. normal-length transcript, longer mRNA, shorter nonfunctional protein e. transcript never madearrow_forwardMicroarrays can be used to determine relative levels of gene expression. In one type of microarray, hybridization of red (experimental) and green (control) labeled cDNAs is proportional to the relative amounts of mRNA in the samples. Red indicates the overexpression of a gene and green indicates the underexpression of a gene in the experimental cells relative to the control cells, yellow indicates equal expression in experimental and control cells, and no color indicates no expression in either experimental or control cells.In one experiment, mRNA from a strain of antibiotic-resistant bacteria (experimental cells) is converted into cDNA and labeled with red fluorescent nucleotides; mRNA from a nonresistant strain of the same bacteria (control cells) is converted into cDNA and labeled with green fluorescent nucleotides. The cDNAs from the resistant and nonresistant cells are mixed and hybridized to a chip containing spots of DNA from genes 1 through 25. The results are shown in the…arrow_forwardMicroarrays can be used to determine relative levels of gene expression. In one type of microarray, hybridization of red (experimental) and green (control) labeled cDNAs is proportional to the relative amounts of mRNA in the samples. Red indicates the overexpression of a gene and green indicates the underexpression of a gene in the experimental cells relative to the control cells, yellow indicates equal expression in experimental and control cells, and no color indicates no expression in either experimental or control cells.arrow_forward
- Researchers have identified a gene (FR) responsible for watermelon resistance to infection by Dacus curcurbitae (a close relative of Drosophila melanogaster). They isolate RNA from resistant (FR+) and sensitive (fr-) watermelons and use a probe that will recognize both FR+ and fr- transcripts. They also isolate protein from resistant and sensitive watermelons and perform a Western blot using an antibody that can recognize the fr- and FR+ protein. Describe the results illustrated below and give a plausible molecular explanation for these observations.arrow_forwardIn the module, you have learned about P-element mediated transgenesis in Drosophila and the concept of using transgenes to rescue mutant phenotypes. In the figure below, you will see a wild type fly with its natural eye colour and three mutants with their eye colours changed to vermillion, white and rosy, respectively. A schematic of P-element mediated transgenesis (as shown in the lectures) is also included in the figure. Please inspect the schematic carefully and choose which of the following statements is true: I. Injection of the white experimental transgene into the vermillion mutant embryo will not change the vermillion mutant phenotype II. Injection of the white experimental transgene in the rosy mutant embryo will change rosy eye colour to red (wild type) III. Injection of the white experimental transgene in the white mutant embryo will not change the white mutant phenotype IV. Injection of the white experimental transgene in the rosy mutant…arrow_forwardIn Figure 13-5, two different methods are illustrated forvisualizing gene expression in developing animals.Which method would allow one to detect where within acell a protein is localized?arrow_forward
- Explain one experimental strategy for determining the functional role of the mouse HoxD-3 gene.arrow_forwardWhen Laybourne and Kadonaga studied the effects of histone proteins on eukaryotic transcription using an in vitro transcription assay explain why: a) they used two different DNA templates that contained different promoter structures. b) when they included both activator protein and histones, they always added the histone proteins before adding the activator to the transcription assay mixture. (Ctri) -arrow_forwardAn electrophoretic mobility shift assay can be used to study the binding of proteins to a segment of DNA. In the results shown here, an EMSA was used to examine the requirements for the binding of RNA polymerase |l (from eukaryotic cells) to the promoter of a protein-encoding gene. The assembly of general transcription factors and RNA polymerase Il at the core promoter is described in Week 4. In this experiment, the segment of DNA containing a promoter sequence was 1100 bp in length. The fragment was mixed with various combinations of proteins and then subjected to an EMSA. Lane 1: No proteins added Lane 2: TFIID Lane 3: TFIIB Lane 4: RNA polymerase IIl Lane 5: TFIID + TFIIB Lane 6: TFIID + RNA 1 2 3 4 5 6. 7 polymerase II Lane 7: TFIID + TFIIB + RNA polymerase Il 1100 bp Explain the results.arrow_forward
- What is the advantage and disadvantage of gene repression in development and What does it mean when we say "enhancers work in a combinatorial fashion?arrow_forwardRecombinant expression in prokaryotic systems has numerous advantages when compared to eukaryotic systems, one of which is the ability to produce the protein of interest at high levels. For this, it is essential to use strong promoters and genetically modified bacteria capable of overexpressing the exogenous gene. Therefore, mark the alternative that best represents the set of bacterial promoters/strains for protein overexpression. * A)use of the T7 promoter, whose induction occurs by the addition of IPTG in the culture medium, and use of the Escherichia coli strain BL21DE3. B)use of the T7 promoter, whose induction occurs by the addition of IPTG in the culture medium, and use of the Escherichia coli DH5 strain. C)use of the lac promoter, whose induction occurs by the addition of IPTG in the culture medium, and use of the Escherichia coli DH5 strain. D) use of the constitutive trp promoter, whose induction occurs by the addition of the amino acid Tryptophan in the medium, and use of…arrow_forwardPROTEIN X HAS THE POTENTIAL FOR MEDICAL APPLICATION IN HUMAN. IT IS ORIGINALLY FROM A PLANT AND HAD BEEN SUCCESSFULLY CLONED FROM ITS GENE. AFTER MANY ATTEMPTS TO CLONE IT INTO EUKARYOTIC EXPRESSION VECTORS WERE UNSUCCESSFUL, THE RESEARCHER DECIDED TO USE E.COLI EXPRESSION VECTOR. HOW THE RECOMBINANT PROTEIN COULD BE EXPRESSED IN E.COLI EXPRESSION VECTOR.1. FACTORS AFFECTING BACTERIAL EXPRESSION SYSTEM.2. STRATEGIES TO IMPROVE THE PROTEIN EXPRESSION.3. WAYS TO ENSURE THE PROTEIN IS ACTIVELY PRODUCED.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
