
Concept explainers
In the following drawing, the top strand is the template DNA, and the bottom strand shows the lagging strand prior to the action of DNA polymerase I. The lagging strand contains three Okazaki fragments. The RNA primers have not yet been removed.
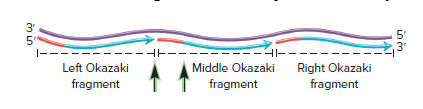
A. Which Okazaki fragment was made first, the one on the left or the one on the right?
B. Which RNA primer will be the first one to be removed by DNA polymerase I, the primer on the left or the primer on the right? For this primer to be removed by DNA polymerase I and for the gap to be filled in, is it necessary for the Okazaki fragment in the middle to have already been synthesized? Explain.
C. Let’s consider how DNA ligase connects the left Okazaki fragment with the middle Okazaki fragment. After DNA polymerase I removes the middle RNA primer and fills in the gap with DNA, where does DNA ligase function? See the arrows on either side of the middle RNA primer. Is ligase needed at the left arrow, at the right arrow, or both?
D. When connecting two Okazaki fragments, DNA ligase uses
or ATP as a source of energy to catalyze this reaction. Explain why DNA ligase needs another source of energy to connect two nucleotides, but DNA polymerase needs nothing more than the incoming
Trending nowThis is a popular solution!

Chapter 11 Solutions
Genetics: Analysis and Principles
- Every tutor here has got this wrong, don't copy off them.arrow_forwardSuppose that the population from question #1 (data is in table below) is experiencing inbreeding depression (F=.25) (and no longer experiencing natural selection). Calculate the new expected genotype frequencies (f) in this population after one round of inbreeding. Please round to 3 decimal places. Genotype Adh Adh Number of Flies 595 Adh Adh 310 Adhs Adhs 95 Total 1000 fladh Adh- flAdn Adh fAdhs Adharrow_forwardWhich of the following best describes why it is difficult to develop antiviral drugs? Explain why. A. antiviral drugs are very difficult to develop andhave no side effects B. viruses are difficult to target because they usethe host cell’s enzymes and ribosomes tometabolize and replicate C. viruses are too small to be targeted by drugs D. viral infections usually clear up on their ownwith no problemsarrow_forward
- This question has 3 parts (A, B, & C), and is under the subject of Nutrition. Thank you!arrow_forwardThey got this question wrong the 2 previous times I uploaded it here, please make sure it's correvct this time.arrow_forwardThis question has multiple parts (A, B & C), and under the subject of Nutrition. Thank you!arrow_forward
- Calculate the CFU/ml of a urine sample if 138 E. coli colonies were counted on a Nutrient Agar Plate when0.5 mls were plated on the NA plate from a 10-9 dilution tube. You must highlight and express your answerin scientific notatioarrow_forwardDon't copy off the other answer if there is anyarrow_forwardAnswerarrow_forward
- HAND DRAW There should be two proarrow_forwardMolecular Biology Question. Please help solve. Thanks. Please draw how two nucleotide triphosphates are linked together to form a dinucleotide, and label the 5' and 3' ends of the resulting dinucleotide.arrow_forwardWhat is a reversion in molecular biology?arrow_forward
 Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning
Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage LearningCase Studies In Health Information ManagementBiologyISBN:9781337676908Author:SCHNERINGPublisher:Cengage
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage LearningCase Studies In Health Information ManagementBiologyISBN:9781337676908Author:SCHNERINGPublisher:Cengage Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax





