
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Textbook Question
Chapter 10, Problem 16P
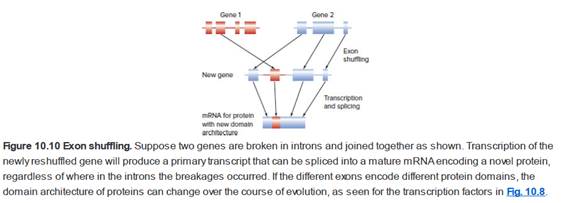
Fig. 10.10 presents a model for exon shuffling in which chromosomal fragments broken into introns can be restitched together to make novel genes that did not exist before. However, only a certain fraction of events like those shown on the figure could actually produce genes encoding proteins with functional domains from both original polypeptides. What is this fraction?
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
You have created three different mutations in the histoneH1 protein (HISmut1, HISmut2, HISmut3), and each of these mutations eliminate a stretch of 5 amino acids from the primary sequence. Based on the description of where you find the mutant histoneH1 proteins when you look inside a cell in each of the cases below, describe 1) what the function is of the amino acids that were removed, and 2) what is not happening with the mutant histoneH1 protein that does happen with wild type H1:
1. HISmut2 protein is found in the cytoplasm, and never in the nucleus.
2. HISmut1 protein is found in only briefly in the cytoplasm because it is very quickly sent to the proteasome.
3. HISmut3 protein is found floating freely throughout the nucleoplasm.
Suppose that the diagram below represents the genomic organization of an enzyme involved in eye pigment production in mice. Within the gene are four exons. Biochemical analysis has revealed that the active site of the enzyme is located in the C terminus of the protein.
-The nucleotide length of each exon and intron is shown.
-The dinucleotide sequence GT represents the 5’ splice site and the dinucleotide sequence AG represents the 3’ splice site. Both the 5’ and the 3’ splice sites must be present for splicing to occur.
Assume that the first and second stop codons are located immediately after the first and second 5’ splice sites, respectively; the third and fourth stop codons are located near the 3’ end of exons 3 and 4, respectively; all these stop codons are in the correct reading frame.
a) draw what the processed mRNA will look like. Include the start codon on the mRNA and label the approximate locations of the 5’ UTR and 3’ UTR on the transcript. (You do not need to add the 5’ CAP…
Shown here is a theoretical viral mRNA sequence 5′-AUGCAUACCUAUGAGACCCUUGGA-3′ (a) Assuming that it could arise from overlapping genes, how many different polypeptide sequences can be produced? Using the chart in Figure 12–7, what are the sequences? (b) A base-substitution mutation that altered the sequence in part (a) eliminated the synthesis of all but one polypeptide. The altered sequence is shown below. Use Figure 12–7 to determine why it was altered. 5′-AUGCAUACCUAUGUGACCCUUGGA-3′
Chapter 10 Solutions
Genetics: From Genes to Genomes
Ch. 10 - Prob. 1PCh. 10 - List three independent techniques you could use to...Ch. 10 - Figure 10.2a has numbers indicating the...Ch. 10 - Which of the enzymes from the following list would...Ch. 10 - Prob. 5PCh. 10 - a. What sequence information about a gene is...Ch. 10 - Why do geneticists studying eukaryotic organisms...Ch. 10 - Consider three different kinds of human libraries:...Ch. 10 - The human genome has been sequenced, but we still...Ch. 10 - This problem investigates issues encountered in...
Ch. 10 - For the sake of simplicity, Fig. 10.4 omitted one...Ch. 10 - Give two different reasons for the much higher...Ch. 10 - Using a cDNA library, you isolated two different...Ch. 10 - The figure that follows shows part of a modified...Ch. 10 - In Problem 14, cDNAs F and G could not be found in...Ch. 10 - Fig. 10.10 presents a model for exon shuffling in...Ch. 10 - An interesting phenomenon found in vertebrate DNA...Ch. 10 - a. If you found a zinc-finger domain which...Ch. 10 - Prob. 19PCh. 10 - In the human immune system, so-called B cells can...Ch. 10 - Chimpanzees have a set of hemoglobin genes very...Ch. 10 - Complete genome sequences indicate that the human...Ch. 10 - On your computers browser, view the page accessed...Ch. 10 - Prob. 24PCh. 10 - Prob. 25PCh. 10 - Certain individuals with mild forms of...Ch. 10 - The 1 and 2 genes in humans are identical in their...Ch. 10 - Prob. 28P
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Given that there are about 20,000 human genes, how can human cells make 75,00o-100,000 different proteins? Distinguish between missense and nonsense. Compare and contrast between insertions and deletions. Why are these called “frameshift” mutations? What are thymine dimers? What causes them?arrow_forwardHuman Chromosome 22 (48 × 106 nucleotide pairs in length) has about 700 protein-coding genes, which average 19,000 nucleotide pairs in length and contain an average of 5.4 exons, each of which averages 266 nucleotide pairs. What fraction of the average protein-coding gene is converted into mRNA? What fraction of the chromosome do these genes occupy?arrow_forwardSuppose that a 20-bp deletion occurs in the middle of exon 2 of the gene depicted in Figure 14.12a. What will be the likely effect of this deletion in the proteins produced by alternative splicing?arrow_forward
- If mature eukaryotic MRNA is hybridized with its corresponding genomic DNA template strand and visualized by electron microscopy, two types of structures are seen: RNA:DNA double-stranded heteroduplexes and single stranded DNA loop structures, as shown in the diagrams below. What do you think these single stranded DNA loops represent? (a) Micrograph of DNA-RNA hybrid (b) Interpretation of micrograph Single-stranded DNA only Single-stranded DNA base paired with MRNA Select one: а. Exons b. Introns c. 5' UTR d. 3' UTR e. promoterarrow_forwardExpression of recombinant proteins in yeast is an important tool for biotechnology companies that produce new drugs for human use. In an attempt to get a new gene X expressed in yeast, a researcher has integrated gene X into the yeast genome near a telomere. Will this strategy result in good expression of gene X? Why or why not? Would the outcome of this experiment differ if the experiment had been performed in a yeast line containing mutations in the H3 or H4 histone tails?arrow_forwardIntrons in protein-coding genes of some eukaryotes are rarely shorter than 65 nucleotides long. What might be a rationale for this limitation?arrow_forward
- Antibiotics such as chloramphenicol, tetracycline, and erythromycin inhibit protein synthesis in bacteria, but have no effect on the synthesis of proteins encoded by eukaryotic nuclear genes. Cycloheximide inhibits the synthesis of proteins encoded by nuclear genes, but has no effect on bacterial protein synthesis. How might these compounds be used to determine which proteins are encoded by mitochondrial and chloroplast genomes?arrow_forwardIn the first figure of “DNMT3L Connects Unmethylated Lysine 4 of Histone H3 to de novo Methylation of DNA,” the authors determined that the DNMT3L protein interacts with several histone proteins. Using Figure a and b (from that paper), explain how each of the 4 histones interact with DNMT3L.arrow_forwardShown below is an R loop prepared for electron microscopy by annealing a purified eukaryotic messenger RNA with DNA from a genomic clone containing the full-length gene corresponding to the mRNA. (a) How many exons does the gene contain? How many introns? (b) Where in this structure would you expect to find a 5′,5′-internucleotide bond? Where would you expect to find a polyadenylic acid sequence?arrow_forward
- In the corn snake Pantherophis guttatus, there are several different color variants, including amelanistic snakes whose skin patterns display only red and yellow pigments. The cause of amelanism in these snakes was recently identified as the insertion of a transposable element into an intron in the OCA2 (oculocutaneous albinism) gene. How might the insertion of extra genetic material into an intron lead to a nonfunctional protein? H▾▾ B 2. = = 66 X² I A▾ X₂ Xarrow_forwardThe following figure represents the primary transcript of a typical eukaryotic protein-coding gene: Polyadenylation site Exon 1 Intron 1 Exon 2 Intron 2 Exon 3 bp: 1 50 600 945 1205 1393 1725 a. What is the size in bases of the fully processed, mature mRNA? Assume a poly (A) tail of 200 adenine residues in your calculations. b. This primary transcript is responsible for the production of a 294 amino acid protein in the liver and a 270 amino acid protein in the brain. Explain how this is possible. 2.arrow_forwardYou would like to add a nuclear localization sequence (NLS) of Lys-Lys-Lys-Arg-Lys to a protein that is usually found in the cytoplasm of a yeast cell. To accomplish this, you introduce the nucleotide sequence encoding the NLS into the gene that encodes the cytoplasmic protein of interest. a. What is the size of the nucleotide insert that will encode the NLS? Briefly explain. 5' 3' b. Below is a diagram of the gene encoding the cytoplasmic protein of interest in the yeast genome. If your goal is to put the NLS at the carboxyl (C) terminus of the protein, at which location (A-E) should the NLS be inserted? Briefly explain. A TATAA ATATT promoter +1 B ATG TAC D TAA ATT stop codon E 3' 5'arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
QCE Biology: Introduction to Gene Expression; Author: Atomi;https://www.youtube.com/watch?v=a7hydUtCIJk;License: Standard YouTube License, CC-BY