
Concept explainers
To analyze:
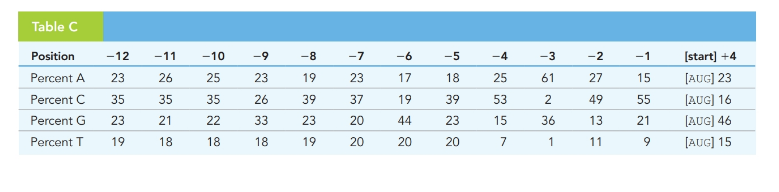
Table-C contains DNA-sequence information compiled by Marilyn Kozak (
Introduction:
Consensus sequences are a short stretch of DNA sequences that are commonly located nucleotides found at a specific location of DNA and RNA. These sequences are used for inter or intramolecular interactions. These sequences are similar in structure and function in all organisms. The consensus sequence is a calculated order that states the most frequent nucleotide at that position in the alignment.
Want to see the full answer?
Check out a sample textbook solution
Chapter 9 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Artificial Selection: Explain how artificial selection is like natural selection and whether the experimental procedure shown in the video could be used to alter other traits. Why are quail eggs useful for this experiment on selection?arrow_forwardDon't give AI generated solution otherwise I will give you downwardarrow_forwardHello, Can tou please help me to develope the next topic (in a esquematic format) please?: Function and Benefits of Compound Microscopes Thank you in advance!arrow_forward
- Identify the AMA CPT assistant that you have chosen. Explain your interpretation of the AMA CPT assistant. Explain how this AMA CPT assistant will help you in the future.arrow_forwardwhat is the difference between drug education programs and drug prevention programsarrow_forwardWhat is the formula of Evolution? Define each item.arrow_forward
- Define the following concepts from Genetic Algorithms: Mutation of an organism and mutation probabilityarrow_forwardFitness 6. The primary theory to explain the evolution of cooperation among relatives is Kin Selection. The graph below shows how Kin Selection theory can be used to explain cooperative displays in male wild turkeys. B When paired, subordinant males increase the reproductive success of their solo, dominant brothers. 0.9 C 0 Dominant Solo EVOLUTION Se, Box 13.2 © 2023 Oxford University Press rB rB-C Direct Indirect Fitness fitness fitness gain Subordinate 19 Fitness After A. H. Krakauer. 2005. Nature 434: 69-72 r = 0.42 Subordinant Dominant a) Use Hamilton's Rule to show how Kin Selection can support the evolution of cooperation in this system. Show the math. (4 b) Assume that the average relatedness among male turkeys in displaying pairs was instead r = 0.10. Could kin selection still explain the cooperative display behavior (show math)? In this case, what alternative explanation could you give for the behavior? (4 pts) 7. In vampire bats (pictured below), group members that have fed…arrow_forwardExamine the following mechanism and classify the role of each labeled species in the table below. Check all the boxes that applyarrow_forward
- 1. Define and explain the two primary evolutionary consequences of interspecific competitionarrow_forward2 A linear fragment of DNA containing the Insulin receptor gene is shown below, where boxes represent exons and lines represent introns. Assume transcription initiates at the leftmost EcoRI site. Sizes in kb are indicated below each segment. Vertical arrows indicate restriction enzyme recognition sites for Xbal and EcoRI in the Insulin receptor gene. Horizontal arrows indicate positions of forward and reverse PCR primers. The Horizontal line indicates sequences in probe A. Probe A EcoRI Xbal t + XbaI + 0.5kb | 0.5 kb | 0.5 kb | 0.5kb | 0.5 kb | 0.5 kb | 1.0 kb EcoRI On the gel below, indicate the patterns of bands expected for each DNA sample Lane 1: EcoRI digest of the insulin receptor gene Lane 2: EcoRI + Xbal digest of the insulin receptor gene Lane 3: Southern blot of the EcoRI + Xbal digest insulin receptor gene probed with probe A Lane 4: PCR of the insulin receptor cDNA using the primers indicated Markers 6 5 4 1 0.5 1 2 3 4arrow_forward4. (10 points) woman. If both disease traits are X-linked recessive what is the probability A man hemizygous for both hemophilia A and color blindness mates with a normal hemophilia A nor colorblindness if the two disease genes show complete that a mating between their children will produce a grandson with neither a. linkage? (5 points) that a mating between their children will produce a grandson with both hemophilia A and colorblindness if the two disease genes map 40 cM apart? (5 points)arrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

