
Concept explainers
QUANTITATIVE More Enzyme Kinetics. The galactose formed in Reaction 6-17 can be phosphorylated by the transfer of a phosphoryl group from ATP, a reaction catalyzed by the enzyme galactokinase:
Assume that you have isolated the galactokinase enzyme and have determined its kinetic parameters by varying the concentration of galactose in the presence of a constant, high (i.e., saturating) concentration of ATP. The double-reciprocal (Lineweaver–Burk) plot of the data is shown as Figure 6-17.
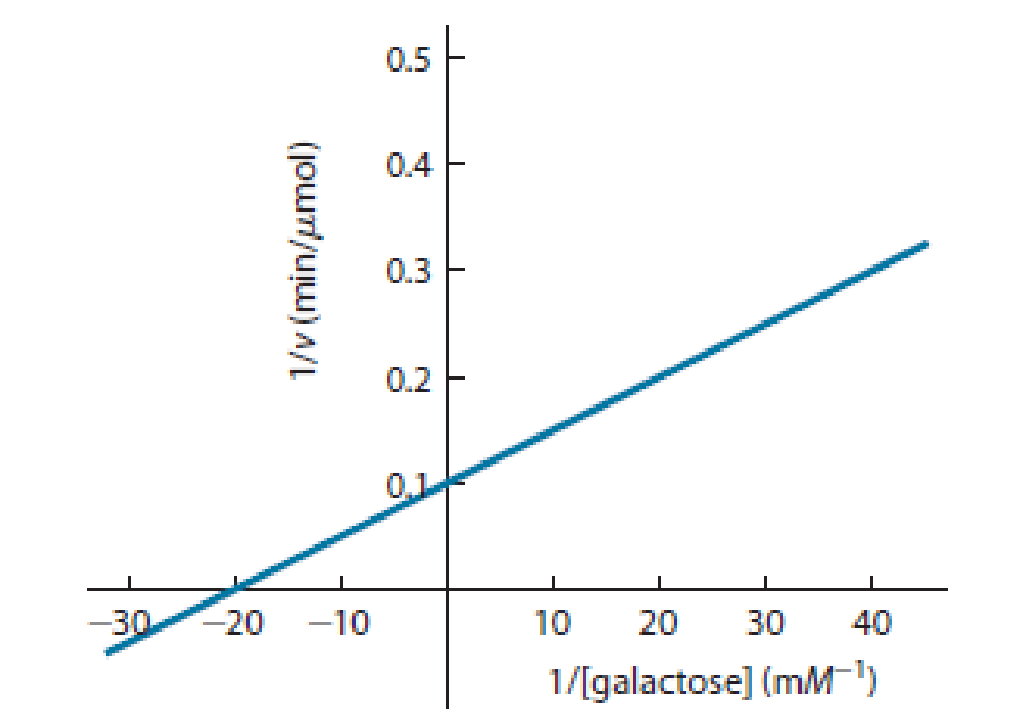
Figure 6-17 Double-Reciprocal Plot for the Enzyme Galactokinase. See Problem 6-7.
(a) What is the Km of galactokinase for galactose under these assay conditions? What does Km tell us about the enzyme?
(b) What is the Vmax of the enzyme under these assay conditions? What does Vmax tell us about the enzyme?
(c) Assume that you now repeat the experiment, but with the ATP concentration varied and galactose present at a constant, high concentration. Assuming that all other conditions aremaintained as before, would you expect to get the same Vmax value as in part b? Why or why not?
(d) In the experiment described in part c, the Km value turned out to be very different from the value determined in part b. Can you explain why?
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Becker's World of the Cell (9th Edition)
- Chemical labeling of chymotrypsin by the compound tosylphenylalanine chloromethyl ketone (TPCK) modifies the His 57 in the enzyme's active site. The structure of this derivative is shown below. TPCK inactivates the enzyme because the bulky addition prevents it from cleaving nearby covalent bonds. HCI + CH, C-O Chymotrypsin-His 57 TPCK Modified enzyme True O Falsearrow_forwardWhat is the catalytic efficiency of Catalase ? Table. The values of KM and kcat for some Enzymes and Substrates Enzyme Carbonic anhydrase Substrate CO2 HCO3 KM (M) 1.2 x 10-2 2.6 x 10-2 Kcat (s-1) 1.0 x 106 4.0 x 105 Catalase H2O2 2.5 x 10-2 1.0 x 107 Urease Urea 2.5 x 10-2 4.0 x 105 O A. 4 x 108 M-s-1 O B. 4 x 108 M-1.s-1 OC25x 10-9 M-s1 D. 2.5 x 102 M-1.s-1 OE 1.0 x 107 s1arrow_forward70 gram lactose working under aerobic conditions. First calculate the total amount of energy units (ATP, GTP, FADH2, NADH) obtained from its degradation to CO2 and H2O. Convert this value to total ATP units.Show your work in detail at each reaction step. Which energy units are obtained after each reaction step.arrow_forward
- ATP yield. Each of the following molecules is processed by glycolysis to lactate. How much ATP is generated from each molecule?arrow_forwardInstructions. Given each set of information which may include common name(s) and the reaction catalyzed, you are required to identify the main class of the specific enzyme described. Name: citryl-CoA synthetase Reaction: ATP + citrate + CoA = ADP + phosphate + (3S)-citryl-CoA Name: D-xylulose reductase Reaction: xylitol + NAD+ = D-xylulose + NADH + H+ Name: cellobiose phosphorylase Reaction: cellobiose phosphate = α-D-glucose 1-phosphate + D-glucose Name: carbonic anhydrase Reaction: H2CO3 = CO2 + H2O Other info: The enzyme catalyzes the reversible hydration of gaseous CO2 to carbonic acid, which dissociates to give hydrogencarbonate above neutral pH. Name: pantoate activating enzyme Reaction: ATP + (R)-pantoate = AMP + diphosphate + (R)-pantothenate.arrow_forwardMATHEMATICAL What yield of ATP can be expected from complete oxidation of each of the following substrates by the reactions of glycolysis, the citric acid cycle, and oxidative phosphorylation? (a) Fructose-1,6-bisphosphate (b) Glucose (c) Phosphoenolpyruvate (d) Glyceraldehyde-3-phosphate (e) NADH (f) Pyruvatearrow_forward
- RECALL Several of the enzymes of glycolysis fall into classes that we will see often in metabolism. What reaction types are catalyzed by each of the following: (a) Kinases (b) Isomerases (c) Aldolases (d) Dehydrogenasesarrow_forwardnot true about the Michaelis-Menten equation? The equation that gives the rate, v, of an the substrate concentration [S] is the Michaelis-Menten equation = Vmax[S]/(Km + [S]), where V, enzyme-catalyzed reaction for all values of max and Km are constants. Which of the following is a) for [S] << Km, V = Vmax applies to most enzymes, but allosteric enzymes have different kinetics when [S] = Km, then v = Vmax/2 gives the rate when the enzyme concentration, temperature, pH, and ionic strength are constant for very high values of [S], v approaches Vmax e) Which is correct about the constant Km in the Michaelis-Menten equation? also called the catalytic constant or turnover number equal to the number of product molecules produced per unit time when the enzyme is saturated with substrate it is the constant in the first order rate equation v = k[A] it is the constant in the second order rate equation v = equal to the substrate concentration at which the velocity or rate of a reaction is ½ the…arrow_forwardNeed help ASAP. Describe the steps by which the F0 portion of the ATP synthase harnesses the proton-motive force to help synthesize ATP. What would you expect to observe if the proton gradient were reversed? Explain your answer.arrow_forward
- Question #1: Choanoflagelletes are a unicellular ancestor to animals. One observation to support this hypothesis is the appearance of adhesion molecules that are key to the development of multiceullarity. Bulk transport Gap junctions Animals. Adhesion, cell signaling Single-celled :} Insects, mammals, and other animals with bilateral symmetry (~10,000,000) Jellyfish and their relatives (10,000) } Sponges (10,000) Choanoflagellates (150) Despite their simple unicellular lifestyle they express adhesion molecules including cadherins and lectins (King et al., 2003) but don't seem to have molecules that are typically found in the extracellular matrix such as integrins or laminins (Williams et al., 2014). Design a microscopy experiment to test the assertion that choanoflagellates have (some) adhesion molecules and those molecules play a similar role in a closely related animal like sponges.arrow_forwardHelp please. This question is specifically asking for the identification of the biomolecules that are attached to the sphingosine core, then we need to answer what bond causes those biomolecules to be connected to the sphingosine, what reaction created that bond (maybe addition or oxidation etc.), what were the starting materials and lastly what reagents or conditions are needed for the reaction to occur. Thank you!arrow_forwardFor 100 words. What are the two essential requirements to effectively carry out metabolic work?arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning

