
Concept explainers
Cartoon renderings of the proteins Top 7 and adaH2 are shown below. Both are soluble, densely packed proteins of roughly 96 residues, and each has a topology of 2 a helices packed onto a 5-stranded ß sheet.
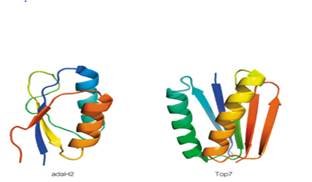
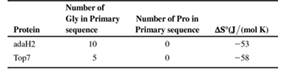
The accompanying table lists some information about the amino acid composition and values of ΔS0 for the folding of these proteins (i.e. for Unfolded →Folded) at 250C. Based on the information in the table, which protein do you predict buries the greater hydrophobic surface area upon folding? Assume 2-state folding (i.e., no intermediates), and that the Unfolded state for both proteins is 100% solvent-exposed. Explain your answer in terms of expected contributions from ΔS0peptide and ΔS0solvent.
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Biochemistry: Concepts and Connections (2nd Edition)
- Do you expect a Pro → Gly mutation in a surface-loop region of a globular protein to be stabilizing or destabilizing? Assume the mutant folds toa native-like conformation. Explain your answer in terms of the predictedenthalpic and entropic effects of the mutation on the ∆G for protein foldingcompared to ∆G of folding for the wild-type proteinarrow_forwardBacteriorhodospin (Mw = 26 kDa) is a protein with purple colour. The protein acts as a light-activated proton pump that provides energy for cellular functions. The structure, revealed by both electron microscopy and X-ray crystallography, showed that the protein consists of 7 transmembrane spanning helices (TMH). Each TMH traverses the lipid bilayer completely (thickness of 45 Å). a)Calculate the minimum number of amino acid residues needed per TMH to completely cross the membrane. b)Estimate how large a fraction of the bacteriorhodopsin protein that is made up by TMHs. c)Now that you know the fraction of the protein that is made up by TMHs, at which ψ/φangles will the amino acids cluster in a Ramachandran plot?arrow_forwardThe process of a protein folding from an inactive unfolded structure to theactive folded structure can be represented by the following equation: unfolded protein ⇌ folded proteinThe values of ΔH° and ΔS° for the folding of the protein lysozyme are: ΔH ° = -280 kJ/ mol ΔS ° = -790 J/mol • K(a) Calculate the value of ΔG° for the folding of lysozyme at 25 °C.(b) At what temperature would you expect the unfolding of lysozyme tobecome favorable? (c) At what temperature would the ratio of unfolded protein to foldedprotein be 1:5?arrow_forward
- Which intermolecular forces are important in acetic acid, CH3 –(C=0)-oh? A particular amino acid contains a- CHNH3+ group. Is this amino acid more likely to be found on the inside or the outside of the folded protein? Briefly explain. The addition of ethanol, CH3CHOH, t an aqueous solution lowers the surface tension of the solution. Predict whether adding ethanol to an aqueous protein solution will tend to stabilize or unfold the protein. Briefly explain.arrow_forwardDo you expect a Pro → Gly mutation in a surface-loop region of a globular protein to be stabilizing or destabilizing? Assume the mutant folds to a native-like conformation. Explain your answer in terms of the predicted enthalpic and entropic effects of the mutation on the AG for protein folding compared to AG of folding for the wild-type protein.arrow_forwardProtein concentration can readily be determined using the Beer-Lambert law: A = e l c where A = absorbance e = molar absorption coefficient (M-1cm-1) l = light path length (cm) c = concentration (M) If the molar absorption coefficient at 280 nm for yeast ADH is 48860 M-1cm-1 and a 10 mL solution of the protein has an absorbance at 280 nm of 0.4 (as measured by a spectrometer with pathlength 1 cm), then what is the concentration of the protein solution (in μM)? i.e. concentration = ______ μM If the molecular weight of the protein is 36849, what is its concentration in mg/mL? i.e. concentration = _______ mg/mL For each part of the question, show your calculations to arrive at your answers.arrow_forward
- The major difference between a protein molecule in its native state and in its denatured state lies in the number of conformations available. To a first ap- proximation, the native, folded state can be thought to have one conforma- tion. The unfolded state can be estimated to have three possible orientations about cach bond between residues. (a) For a protein of 100 residues, estimate the entropy change per mole upon denaturation. (b) What must be the enthalpy change accompanying denaturation to allow the protein to be half-denatured at 50 °C? (c) Will the fraction denatured increase or decrease with increasing temperature?arrow_forwardTERTIARY STRUCTURE (A) (B) (C) Fg Eet Galand Sen 20e Figure 6. Examples of the arrangement of a-helices and B-sheets in folded protein domains. Copyright 2013 from Essential Cell Biology, 4th Edition by Alberts et al. Reproduced by permission of Garland Science/ Taylor & Francis LLC. Figure 6 shows three examples of how secondary structure elements can be arranged in relation to one another in the functional, folded form of a complete protein or one compact portion of a protein. The overall three-dimensional shape (or conformation) of a protein is its tertiary structure. • What do you think holds together the various secondary structural elements in a particular three-dimensional pattern? (Hint: Look back at Figure 5 - what is sticking out from the sides of the a-helices and B-strands?)arrow_forwardLoop regions play important roles in the secondary structure of protein. Define loop region and give three (3) of the rolesarrow_forward
- What would you predict about the ratio of hydrophilic to hydro- phobic amino acid residues in a series of monomeric globular proteins that range in size from a molecular weight of 10,000 g/mole to a molecular weight of 100,000 g/mole? Note that the volume of a sphere is 4/3nr³, while the sur- face area of the outside of a sphere is 4пr².arrow_forwardConsider a protein with two surface-exposed histidine residues: HisA is a “typical” histidine residue with a pKa = 6.2 HisB is involved in a stabilizing interaction (His-NH+ ..... -O2C-Glu) with a neighboring glutamic acid residue. For HisB, the Gibbs free energy of deprotonation at pH = 7.0 and T = 293K is ΔG'o = +15 kj mol-1. If you had a solution, at pH = 7.0 and T = 293K, containing this protein: a) What fraction of HisA residues are protonated? b) What fraction of HisB residues are protonated? c) What is the pKa of HisB?arrow_forwardWhat would you predict about the ratio of hydrophilic to hydro- phobic amino acid residues in a series of monomeric globular proteins that range in size from a molecular weight of 10,000 g/mole to a molecular weight of 100,000 g/mole? Note that the volume of a sphere is 4/3nr³, while the sur- face area of the outside of a sphere is 4nr². How would your prediction change if the monomeric proteins were fibrous instead of globular?arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





