
Biochemistry: Concepts and Connections (2nd Edition)
2nd Edition
ISBN: 9780134641621
Author: Dean R. Appling, Spencer J. Anthony-Cahill, Christopher K. Mathews
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 6, Problem 13P
A protein gives under conditions of buffer composition, pH, and temperature that are close to physiological conditions, a molecular weight by size exclusion measurements of 140,000 g/mol. When the same protein studied by SDS gel electrophoresis in the absence or presence of the reducing agent
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
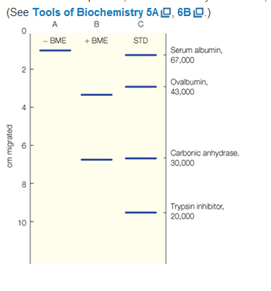
A protein gives, under conditions of buffer composition, pH, and temperature that are close to physiological conditions, a molecular weight by sedimentation equilibrium measurements of 140,000 g/mol. When the same protein is studied by SDS gel electrophoresis in the absence or presence of the reducing agent β-mercaptoethanol (BME), the patterns seen in lanes A and B respectively are observed. Lane C contains standards of molecular weight indicated. From these data, describe the native protein, in terms of the kinds of subunits present, the stoichiometry of subunits (that is, how many of each subunits are present), and the kinds of bonding (covalent, non-covalent) existing between subunits
. A protein gives, under conditions of buffer composition, pH, and tem-
perature that are close to physiological conditions, a molecular weight
by sedimentation equilibrium measurements of 140,000 g/mol. When
the same protein is studied by SDS gel electrophoresis in the absence
or presence of the reducing agent B-mercaptoethanol (BME), the pat-
terns seen, respectively, in lanes A and B are observed. Lane C contains
standards of molecular weight indicated. From these data, describe the
native protein, in terms of the kinds of subunits present, the stoi-
chiometry of subunits, and the kinds of bonding (covalent, noncova-
lent) existing between subunits. /
A
B
- BME
+ BME
STD
Serum albumin,
67,000
Ovalbumin,
43,000
Carbonic anhydrase,
30,000
Trypsin inhibitor,
20,000
10
2.
4.
Co
cm migrated
suctose density gradient ultractifugation is a powerful technique for fractionating macromolecules like DNA,RNA and proteins.some protocols using sucrose gradients mention the following:10 mL sucrose gradients 10-15%(w/v) in 10 mM HEPES buffer.How would you prepare this sucros gradient?
Chapter 6 Solutions
Biochemistry: Concepts and Connections (2nd Edition)
Ch. 6 - Prob. 1PCh. 6 - Bovine pancreatic trypsin inhibitor (BPTI; Figure...Ch. 6 - A schematic structure of the subunit of...Ch. 6 - In the protein adenylate kinase, the C-terminal...Ch. 6 - Give two reasons to explain why a proline residue...Ch. 6 - Consider a small protein containing 101 amino acid...Ch. 6 - a. Based on a more conservative answer to Problem...Ch. 6 - The following sequence is part of a globular...Ch. 6 - a. A protein is found to be a tetramer of...Ch. 6 - Under physiological conditions, the protein...
Ch. 6 - Theoretical and experimental measurements show...Ch. 6 - The peptide hormone vasopressin is used in the...Ch. 6 - A protein gives under conditions of buffer...Ch. 6 - A protein gives a single band on SDS get...Ch. 6 - It has been postulated that the normal...Ch. 6 - Below are shown two views of the backbone...Ch. 6 - Do you expect a Pro Gly mutation in a...Ch. 6 - Rank the following in terms of predicted rates...Ch. 6 - Shown below are two cartoon views of the small...Ch. 6 - Prob. 20PCh. 6 - In most cases, mutations in the core of protein...Ch. 6 - A Leu Ala mutation at a site buried the core of...Ch. 6 - Disulfide bonds have been shown to stabilize...Ch. 6 - Cartoon renderings of the proteins Top 7 and adaH2...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biochemistry and related others by exploring similar questions and additional content below.Similar questions
- From this standard curve and chart below, does the separation of molecules in the mixture appear successful from the gel filtration? Is there a clearlydefined separation between molecules? Explain your conclusions. Parameters required for calculation of coefficient (Kd) for unknown protein Volume eluted (mL) Which variable does this volume represent in the equation for Kd? Fraction with maximal DNP-Aspartate detected 36 Vt Fraction with maximal Protein detected 24 Ve Fraction with maximal Blue dextran detected 6 Voarrow_forwardA series of standard proteins and an unknown enzyme were studied by gel filtration on a Sephadex G200 (the 200 refers to the maximum pore size in kDa) column. The elution volume Vel for each protein is given in Table 1 below. (a) Plot the data in the form of log Mr versus elution volume. From the line of best fit through the points for the standards, determine the Mr of the unknown enzyme. Explain why ferritin and ovomucoid behave anomalously. Table 1 - The Vel versus Mr data Protein Mr Vel (mL) Blue dextran* Lysozyme Chymotrypsin Ovalbumin Serum albumin Aldolase Urease Ferritin# Ovomucoid# Unknown 1,000,000 14,000 25,000 45,000 68,500 150,000 500,000 700,000 28,000 ⎯ 85 200 190 170 150 125 90 92 160 139 *Blue dextran is not a protein but a high-Mr carbohydrate that has a covalently bound blue dye, and it elutes with the void volume of the column. # Do not…arrow_forwardIn a mixture of five proteins listed, draw an elution profile (Absorbance vs. mL eluted, arbitrary) for the purification of the listed proteins on a gel filtration chromatography resin: cytochrome c (pI = 5.4; Mr = 13,000), immunoglobulin G (pI = 7.3; Mr = 145,000), ribonuclease A (pI = 9.6; Mr = 13,700), RNA polymerase (pI = 6.3; Mr = 450,000), human serum albumin (pI = 5.4; Mr = 68,500). Label your elution peaks. Draw a sketch of an SDS-PAGE, reflecting the mobility of the above mixture as they elute from the column. Label you protein bands.arrow_forward
- The A280 of a protein sample loaded onto a gel was determined to be 0.767 (1.00 cm path length, after subtracting the blank). The total volume of this sample was 428 µL. 19.0 µL of this protein sample was mixed with 19.0 µL of 2X laemalli sample buffer and then 12.0 µL of the entire sample was loaded into the gel and electrophoresed. Calculate the amount of protein that was loaded into the gel (in µg).arrow_forwardThe protein concentration of a known standard is 100mg/mL If you prepared a serial dilution, mixing 10μL of protein with 40μL of water what would be concentrations of the first 3 dilutions?arrow_forwardcalculate the volume of stock solutions required to make up the buffer solutions that will be used for protein purification. The solutions you need to prepare for purification are: i. Binding Solution A: make up 50 mL 50 mM HEPES buffer (pH 7.5), 300 mM NaCl, 5mM imidazole, 5% (v/v) glycerol ii. Wash Solution B: make up 50 mL 50 mM HEPES buffer (pH 7.5), 300 mM NaCl, 75mM imidazole, 5% (v/v) glycerol iii. Elution Solution C: make up 10 mL 50 mM HEPES buffer (pH 7.5), 300 mM NaCl, 500 mM imidazole, 5% (v/v) glycerol please show your working . Thnk youarrow_forward
- 80mL of a 0.3M solution of hexapeptide Leu-His-Cys-Glu-Asn-Arg is adjusted to pH=pl. The solution is then titrated with 0.2M HCI to a final pH of 2.1. Sketch the titration curve, labelling the pH and volume axes. Indicate the volume of HCl needed to reach each relevant pKa value and equivalence point(s). Relevant pka values are: 2.1, 4.3, 6.0, 8.3, 9.8, and 12.5.arrow_forwardProtein concentration can readily be determined using the Beer-Lambert law: A = e l c where A = absorbance e = molar absorption coefficient (M-1cm-1) l = light path length (cm) c = concentration (M) If the molar absorption coefficient at 280 nm for yeast ADH is 48860 M-1cm-1 and a 10 mL solution of the protein has an absorbance at 280 nm of 0.4 (as measured by a spectrometer with pathlength 1 cm), then what is the concentration of the protein solution (in μM)? i.e. concentration = ______ μM If the molecular weight of the protein is 36849, what is its concentration in mg/mL? i.e. concentration = _______ mg/mL For each part of the question, show your calculations to arrive at your answers.arrow_forwardThe diffusivity of amino acids in polyacrylamide gel is approximately 1x10^-9 cm2./s calculate the initial flux of amino acids, give an instantaneous gradient of (20g/cm 3 )/8cm Why is polyacrylamide gel is used in electrophoresis?arrow_forward
- Consider the following properties of the protein components of a sample mixture as provided in the table below: 1. if the mixture is subjected to gel filtration chromotography which protein component elute first? 2. if the mixture is subjected to isoelectric focusing which protein will stop m oving nearest to the positive electrode? 3. if the mixture is subjected to cation-exchange chromotography using a buffer at ph 7 which protein will bind to the resin? 4.if the mixture is subjected to SDS-PAGE which protein will be at bottomost portion of gel? 5.if the mixture is subjected to hydrophobic interaction chromotography which protein will bind most strongly to the resin?arrow_forwardIn RP HPLC, one of the most common separation methods used to measure purity, strength, dosage, etc, a protein would be put into 0.1% (1000 ppm) TFA (Trifluoroacetic acid), what do you suppose this does to the protein in many cases? (Pick the BEST answer). somewhat denaturing to very denaturing oxidizes cysteine ionizes acid and base groupsarrow_forwardDiethylaminoethyl cellulose is a positively charged resin used in ion-exchange chromatography with a pKa is about 11. A negatively charged protein with an isoelectric point of 5.0 is applied to the column in a buffer at pH 7 and containing 0.1 M NaCl. Which of the following conditions is most likely to weaken the interaction between the protein and the resin? A Raising the pH to 8 and decreasing the NaCl to 0.05 M B Raising the pH to 8 and increasing the NaCl to 0.2 M C Lowering the pH to 6 and decreasing the NaCl to 0.05 M D Lowering the pH to 6 and increasing the NaCl to 0.2 Marrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON

Biochemistry
Biochemistry
ISBN:9781319114671
Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.
Publisher:W. H. Freeman

Lehninger Principles of Biochemistry
Biochemistry
ISBN:9781464126116
Author:David L. Nelson, Michael M. Cox
Publisher:W. H. Freeman

Fundamentals of Biochemistry: Life at the Molecul...
Biochemistry
ISBN:9781118918401
Author:Donald Voet, Judith G. Voet, Charlotte W. Pratt
Publisher:WILEY

Biochemistry
Biochemistry
ISBN:9781305961135
Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougal
Publisher:Cengage Learning

Biochemistry
Biochemistry
ISBN:9781305577206
Author:Reginald H. Garrett, Charles M. Grisham
Publisher:Cengage Learning

Fundamentals of General, Organic, and Biological ...
Biochemistry
ISBN:9780134015187
Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. Peterson
Publisher:PEARSON
Biomolecules - Protein - Amino acids; Author: Tutorials Point (India) Ltd.;https://www.youtube.com/watch?v=ySNVPDHJ0ek;License: Standard YouTube License, CC-BY