
Two important methods for understanding the genetic basis for development are mitotic crossing-over and the use of the gene from jellyfish called GFP (for green fluorescent protein) that makes these animals glow in the dark. By recombinant DNA techniques described later in the book, you can insert the jellyfish GFP gene anywhere into the genome of organisms like Drosophila or mice. Cells expressing this GFP gene will glow green in the microscope, while those without the GFP gene will not glow green.
Mice homozygous for the recessive mutation small cells (smc) die as early embryos because their cells divide prematurely before they reach normal size.
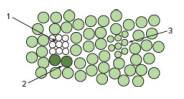
You want to design a mouse carrying one copy of the GFP gene and heterozygous for smc in which you could generate clones in adult mice by mitotic recombination. In this designer mouse, every cell in every clone that is not green would be homozygous for the smc mutation. The figure below shows a field of epithelial cells in the mouse you design. You will see some cells that are normal size and other cells that are small. You will also see cells of three different colors: blank, weakly glowing cells (light green), and brightly glowing cells (dark green). Most of the cells in the epithelium of this mouse are of normal size and weakly glowing. The epithelium also contains three clones of cells (1, 2, and 3) that have unusual appearances due to the occurrence of mitotic recombination.
| a. | Show the chromosomes and centromeres, the alleles smc+ and smc, and GFP+ (GFP gene present) and GFP- (GFP gene absent) in your designer mouse. (As a reminder, this mouse will carry one copy of the GFP gene and will be heterozygous for smc. Every cell in every clone generated by mitotic recombination that is not green should be homozygous for the smc mutation.) |
| b. | Why do you need to use mitotic recombination to study the function of smc+ in adult mice? |
| c. | Why do you see cells of three different colors? |
| d. | Why are clones 1 and 2 next to each other? |
| e. | On your map in part (a), place an arrow to show the position of a mitotic recombination event that could give rise to clones 1 and 2.. |
| f. | Why do more cells exist in clone 1 than in clone 2? |
| g. | On your map in part (a), place an arrow to show the position of a mitotic crossover that could give rise to clone 3. |
Want to see the full answer?
Check out a sample textbook solution
Chapter 5 Solutions
Genetics: From Genes to Genomes
- Northern blotting, RT-PCR, and microarrays can be used to analyze gene expression. A lab studies yeast cells, comparing their growth in two different sugars, glucose and galactose. One student is comparing expression of the gene HMG2 under these two conditions. Which technique(s) could he use and why? Another student wants to compare expression of all the genes on chromosome 4, of which there are approximately 800. What technique(s) could she use and why?arrow_forwardThe GAL4 expression system can be used to experimentally induce expression of a gene in a specific tissue at a specific time. You want to study the overexpression of Your Favorite Gene (YFG) in mouse ear development. However, if you overexpress this gene everywhere, the mouse is not viable past early embryo development. You have the following mouse lines with different mutations or transgenic constructs available. Which two will you cross together to answer your research questions? GAL4 with constitutive enhancer GAL4 with embryo enhancer GAL4 with ear enhancer yfg loss-of-function line YFG CDNA with UASG promoter YFG CDNA with constitutive promoterarrow_forwardOur understanding of maternal effect genes has been greatly aided by their identification in experimental organisms such as Drosophila melanogaster and Caenorhabditis elegans. In experimental organisms with a short generation time, geneticists have successfully searched for mutant alleles that prevent the normal process of embryonic development. In many cases, the offspring die at early embryonic or larval stages. These are called maternal effect lethal alleles. How would a researcher identify a mutation that produced a recessive maternal effect lethal allele?arrow_forward
- Draw a basket mutant embryo. What does basket encode? Why do the mutant embryos have this phenotype?arrow_forwardAnother way to study the role of proteins (e.g., transcription factors) that function in development is to microinject the mRNA that encodes a protein, or the purified protein itself, into an oocyte or embryo, and then determine how this affects the subsequent development of the embryo, larva, and adult. For example, if Bicoid protein is injected into the posterior region of an oocyte, the resulting embryo will develop into a larva that has anterior structures at both ends. Based on your understanding of the function of each developmental gene, what would be the predicted phenotype if the following proteins or mRNAs were injected into normal oocytes? A. Nanos mRNA injected into the anterior end of an oocyte B. Antp protein injected into the posterior end of an embryo C. Toll mRNA injected into the dorsal side of an early embryoarrow_forwardA panel of cell lines was created by human–mouse somatic-cell hybridization. Each cell line was examined for the presence of human chromosomes and for the production of three enzymes. The following results were obtained:On the basis of these results, give the chromosomal locations of the genes encoding enzyme 1, enzyme 2, and enzyme 3.arrow_forward
- In contrast with the genomic manipulations of animals and plants described in this chapter, human genetherapy is directed specifically at altering the genomes of somatic cells rather than germ-line cells.Why couldn’t or wouldn’t medical scientists try to alter the genome of human germ-line cells?arrow_forwardScientists carried out a microarray analysis to compare the gene expression of normal pancreatic cells to that of cancer cells from a person with pancreatic cancer. The scientists labeled the cDNA from the normal pancreatic cells with green fluorescent nucleotides. They labeled the cDNA from the cancer cells with red fluorescent nucleotides. The two cDNAs were mixed and allowed to hybridize to a microarray. Less p53 activity is found in cancer pancreatic cells than normal cells. What color would the spot for the p53 gene be on the microarray? Red Green Yellow Blackarrow_forwardNot all inherited traits are determined by nuclear genes (i.e., genes located in the cell nucleus) that are expressed during the life of an individual. In particular, maternal effect genes and mitochondrial DNA are notable exceptions. With these ideas in mind, let’s consider the cloning of a sheep (e.g., Dolly). A. With regard to maternal effect genes, is the phenotype of such a cloned animal determined by the animal that donated the enucleatedegg or by the animal that donated the somatic cell nucleus? Explain.arrow_forward
- A normal appearing female infant was identified with a positive newborn screen, linked to chromosome 12. Few years ago, her older sibling had developed profound hypoglycemia, liver failure leading to coma, and subsequent irreparable brain damage, following a viral illness. The sibling was subsequently shown by clinical testing to have the same disorder that this female infant is screened positive for. a. What is the most likely diagnosis? b. What biomarkers would confirm this on the newborn screening process? Describe the mechanism that causes this metabolic defect.arrow_forwardThe oncogenic protein BETA promotes entry into the S phase of the cell cycle. Phosphorylation of BETA at the amino acid Tyr98 causes BETA to be degraded by the proteasome, thus limiting its abundance. A mutation in the codon encoding Tyr98 changes this residue to Cys, which cannot be phosphorylated. What is the best description of this mutant allele?a) antimorphb) hypermorphc) hypomorphd) amorphe) neomorpharrow_forwardUsing a transgenic technique, propose an experiment to determine whether Cdx2 is sufficient for trophoblast development in the mouse embryo. Describe two results that you would expect to observe at the blastocyst stage if Cdx2 is indeed sufficient for trophoblast development. Be as specific as possible regarding the transgene that you propose for this experiment (including what gene's enhancer you would use in the transgene). Note: you do not need to explain the details of how a transgenic mouse is made. Describe the experiment in steps (Step 1: ..., Step 2: ... etc) and please keep your answer to under 150 words. tips: DONT talk about stop cassetes/memory cassetes, focus on transgenes Paper called "Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst" gave lots of results that you might see,, 6 diff ways that cdx2 is required for trophoblasts need specific gene enhancer (dont just say "expressed enhancer in genital…arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





