
Concept explainers
Interpretation:
The full name of dUMP, UMP, CDP, AMP, and ATP has to be written.
Concept Introduction:
Composition of
Sugar: In both DNA and RNA, sugar portion is found. In DNA, the sugar is D-ribose, where at 2’hydroxyl group is absent and in RNA, the hydroxyl group is present at 2’.
Nitrogenous bases: Five types of nitrogenous bases (has unique one-letter code A, G, T, U, and C) are derived from two parent compounds called purine and pyrimidine. The purine derivatives are Adenine and Guanine are two fused nitrogen containing rings. The pyrimidine derivatives are Thymine, Cytosine, and Uracil are only one nitrogen containing six-membered ring. Adenine, Guanine, Thymine, and Cytosine are the nitrogenous bases present in DNA. Adenine, Guanine, Cytosine and Uracil are the nitrogenous bases present in RNA.
Nucleotide: (Nucleoside + phosphate)
Nucleotides are the building blocks of nuclei acids; monomers of DNA and RNA
Nucleoside and its naming: The combination of monosaccharide (sugar) and nitrogenous base is known as nucleoside. The nucleoside names are the nitrogenous base name modified with criteria. While naming nucleoside of purine derivatives the suffix ‘-osine’ is included and for pyrimidine derivatives the suffix ‘-idine’ is used. No prefix used for the nucleosides containing ribose and the prefix ‘deoxy-’ is used for deoxyribose.
Naming nucleotide: At the end of the nucleoside, phosphate group is added. For example, 5’-monophosphate means adding one phosphate group at 5’carbon in the sugar ring. Uridine monophosphate can be written as UMP.
Numbering the atoms in sugar and base rings:
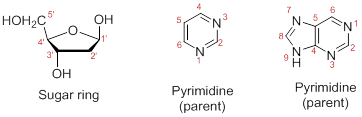
In order to distinguish the atoms in the sugar of a nucleoside and atoms of a base ring, numbers without prime is used for atoms in the base ring and numbers with prime used for the atoms in the sugar ring.
Want to see the full answer?
Check out a sample textbook solution
Chapter 26 Solutions
Fundamentals of General, Organic, and Biological Chemistry (8th Edition)
- Please provide a chromatography technique to isolate the protein A from the mixture containing protein A and B (shown as below). And explain the separation mechanism related to this example. Protein A: MKRHRRKKHHRKRRKKRKGH (positively charged) Protein B: MDEEEEDDDEEDDEEDEDEED (negatively charged)arrow_forwardGive the name of two Molecules of Farnesyl Pyrophosphate:arrow_forwardDraw the GC base pair at pH 10.arrow_forward
- Denatured protein is in a low energy state. What sort of explanation can you use to rationalize that statement? Hint: consider Gibbs free energy.arrow_forwardThe manufacture of chocolates containing a liquid center is an interesting application of enzyme engineering. The flavored liquid center consists largely of an aqueous solution of sugars rich in fructose to providesweetness. The technical dilemma is the following: the chocolate coating must be prepared by pouring hot melted chocolate over a solid (or almost solid) core, yet the final product must have a liquid, fructose-rich center. Suggest a way to solve this problem. (Hint: Sucrose is much less soluble than a mixture of glucose and fructose.)arrow_forwardDraw the following oligonucleotides: a. d(pAACGTCTC) b. pGUAACUGCarrow_forward
- Given the Ramachandran Plot below, identify the protein components that could adopt the phi-psi angle combination indicated by the number 3. (IF YOU COPY UR ANSWER FROM ANOTHER WEBSITE I WILL DISLIKE)arrow_forwardWhich nuclear isotope used in protein NMR spectroscopy is the most sensitive to detect? Briefly explain why.arrow_forwardDiscuss the behavior of enzymes as described by the Michaelis-Menten Equationarrow_forward
- What is Transpeptidation reaction?arrow_forwardResearch the IUPAC and common names and the short-hand codes of the following: 9 saturated fatty acids (4C-20C atoms) 5 unsaturated fatty acids (16C-20C atoms) 3 essential fatty acids 4 eicosanoidsarrow_forwardDuring the early stages of an enzyme purification protocol, when cells have been lysed but cytosolic components have not been separated, the reaction velocity-versus-substrate concentration is sigmoidal. As you continue to purify the enzyme, the curve shifts to the right. Explain your results. This is an allosteric enzyme and you must use a Lineweaver-Burk plot to determine KM and Vmax correctly. This is an enzyme that displays Michaelis-Menten kinetics and you purify away an inhibitor. This is an allosteric enzyme and during purification you purify away an activator. This is an allosteric enzyme displaying a double-displacement mechanism and during purification you purify away one of the substrates: This is an enzyme that displays Michaelis-Menten kinetics, and you must use a Lineweaver-Burk plot to determine KM and Vmax correctly.arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





