
Genetics: Analysis and Principles
6th Edition
ISBN: 9781259616020
Author: Robert J. Brooker Professor Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 23, Problem 17EQ
Four cosmid clones, which we will call cosmids A, B, C, and D, were hybridized to each other in pairwise combinations. The insert size of each cosmid was also analyzed. The following results were obtained:
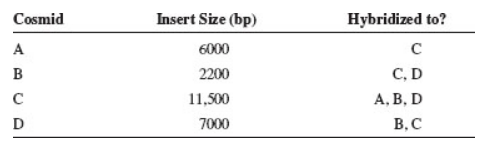
Draw a map that shows the order of the inserts within these four cosmids.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
What did the Cre-lox system used in the Kikuchi et al. 2010 heart regeneration experiment allow researchers to investigate?
What was the purpose of the cmlc2 promoter?
What is CreER and why was it used in this experiment?
If constitutively active Cre was driven by the cmlc2 promoter, rather than an inducible CreER system, what color would you expect new cardiomyocytes in the regenerated area to be no matter what? Why?
What kind of organ size regulation is occurring when you graft multiple organs into a mouse and the graft weight stays the same?
What is the concept "calories consumed must equal calories burned" in regrads to nutrition?
Chapter 23 Solutions
Genetics: Analysis and Principles
Ch. 23.1 - Prob. 1COMQCh. 23.2 - Prob. 1COMQCh. 23.3 - A molecular marker is a _____ found at a specific...Ch. 23.3 - 2. Which of the following is an example of a...Ch. 23.3 - To map the distance between molecular markers via...Ch. 23.4 - 1. What is a contig?
a. A fragment of DNA that...Ch. 23.4 - A vector that can carry a large fragment of...Ch. 23.4 - 3. Chromosomal walking is a method of _____ in...Ch. 23.5 - Prob. 1COMQCh. 23.5 - Prob. 2COMQ
Ch. 23.5 - 3. A prokaryotic genome is about 4 million bp in...Ch. 23.6 - Metagenomics is aimed at a. determining the...Ch. 23 - 1. A person with a rare genetic disease has a...Ch. 23 - For each of the following, decide if it could be...Ch. 23 - Which of the following statements about molecular...Ch. 23 - 1. Is each of the following a method used in...Ch. 23 - Prob. 2EQCh. 23 - Prob. 3EQCh. 23 - The cells from a persons malignant tumor were...Ch. 23 - 5. Figure 23.2 describes the technique of FISH....Ch. 23 - Explain how DNA probes with different fluorescence...Ch. 23 - 7. A researcher is interested in a gene found on...Ch. 23 - Prob. 8EQCh. 23 - Prob. 9EQCh. 23 - Prob. 10EQCh. 23 - Prob. 11EQCh. 23 - Prob. 12EQCh. 23 - In the Human Genome Project, researchers have...Ch. 23 - 14. Take a look at question 3 in More Genetic...Ch. 23 - 15. Place the following stages of a physical...Ch. 23 - 16. What is an STS? How are STSs generated...Ch. 23 - 17. Four cosmid clones, which we will call cosmids...Ch. 23 - A human gene, which we will call geneX, is located...Ch. 23 - 19. Describe how you would clone a gene by...Ch. 23 - 20. A bacterium has a genome size of 4.4 Mb. If a...Ch. 23 - 21. Discuss the advantages of next-generation...Ch. 23 - Prob. 22EQCh. 23 - Prob. 23EQCh. 23 - What is a molecular marker? Give two examples....Ch. 23 - Which goals of the Human Genome Project do you...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- You intend to insert patched dominant negative DNA into the left half of the neural tube of a chick. 1) Which side of the neural tube would you put the positive electrode to ensure that the DNA ends up on the left side? 2) What would be the internal (within the embryo) control for this experiment? 3) How can you be sure that the electroporation method itself is not impacting the embryo? 4) What would you do to ensure that the electroporation is working? How can you tell?arrow_forwardDescribe a method to document the diffusion path and gradient of Sonic Hedgehog through the chicken embryo. If modifying the protein, what is one thing you have to consider in regards to maintaining the protein’s function?arrow_forwardThe following table is from Kumar et. al. Highly Selective Dopamine D3 Receptor (DR) Antagonists and Partial Agonists Based on Eticlopride and the D3R Crystal Structure: New Leads for Opioid Dependence Treatment. J. Med Chem 2016.arrow_forward
- The following figure is from Caterina et al. The capsaicin receptor: a heat activated ion channel in the pain pathway. Nature, 1997. Black boxes indicate capsaicin, white circles indicate resinferatoxin. You are a chef in a fancy new science-themed restaurant. You have a recipe that calls for 1 teaspoon of resinferatoxin, but you feel uncomfortable serving foods with "toxins" in them. How much capsaicin could you substitute instead?arrow_forwardWhat protein is necessary for packaging acetylcholine into synaptic vesicles?arrow_forward1. Match each vocabulary term to its best descriptor A. affinity B. efficacy C. inert D. mimic E. how drugs move through body F. how drugs bind Kd Bmax Agonist Antagonist Pharmacokinetics Pharmacodynamicsarrow_forward
- 50 mg dose of a drug is given orally to a patient. The bioavailability of the drug is 0.2. What is the volume of distribution of the drug if the plasma concentration is 1 mg/L? Be sure to provide units.arrow_forwardDetermine Kd and Bmax from the following Scatchard plot. Make sure to include units.arrow_forwardChoose a catecholamine neurotransmitter and describe/draw the components of the synapse important for its signaling including synthesis, packaging into vesicles, receptors, transporters/degradative enzymes. Describe 2 drugs that can act on this system.arrow_forward
- The following figure is from Caterina et al. The capsaicin receptor: a heat activated ion channel in the pain pathway. Nature, 1997. Black boxes indicate capsaicin, white circles indicate resinferatoxin. a) Which has a higher potency? b) Which is has a higher efficacy? c) What is the approximate Kd of capsaicin in uM? (you can round to the nearest power of 10)arrow_forwardWhat is the rate-limiting-step for serotonin synthesis?arrow_forwardWhat enzyme is necessary for synthesis of all of the monoamines?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Principles Of Radiographic Imaging: An Art And A ...Health & NutritionISBN:9781337711067Author:Richard R. Carlton, Arlene M. Adler, Vesna BalacPublisher:Cengage Learning
Principles Of Radiographic Imaging: An Art And A ...Health & NutritionISBN:9781337711067Author:Richard R. Carlton, Arlene M. Adler, Vesna BalacPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning- Case Studies In Health Information ManagementBiologyISBN:9781337676908Author:SCHNERINGPublisher:Cengage
 Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning
Human Biology (MindTap Course List)BiologyISBN:9781305112100Author:Cecie Starr, Beverly McMillanPublisher:Cengage Learning

Principles Of Radiographic Imaging: An Art And A ...
Health & Nutrition
ISBN:9781337711067
Author:Richard R. Carlton, Arlene M. Adler, Vesna Balac
Publisher:Cengage Learning


Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning

Case Studies In Health Information Management
Biology
ISBN:9781337676908
Author:SCHNERING
Publisher:Cengage

Human Biology (MindTap Course List)
Biology
ISBN:9781305112100
Author:Cecie Starr, Beverly McMillan
Publisher:Cengage Learning

Mitochondrial mutations; Author: Useful Genetics;https://www.youtube.com/watch?v=GvgXe-3RJeU;License: CC-BY