
Concept explainers
QUANTITATIVE Introns. To investigate the possible presence of introns in three newly discovered genes (X, Y, and Z), you perform an experiment in which the restriction enzyme HaeIII is used to cleave either the DNA of each gene or the cDNA made by copying its mRNA with reverse transcriptase. The resulting DNA fragments are separated by gel electrophoresis, and the presence of fragments in the gels is detected by hybridizing to a radioactive DNA probe made by copying the intact gene with DNA polymerase in the presence of radioactive substrates. The following results are obtained:
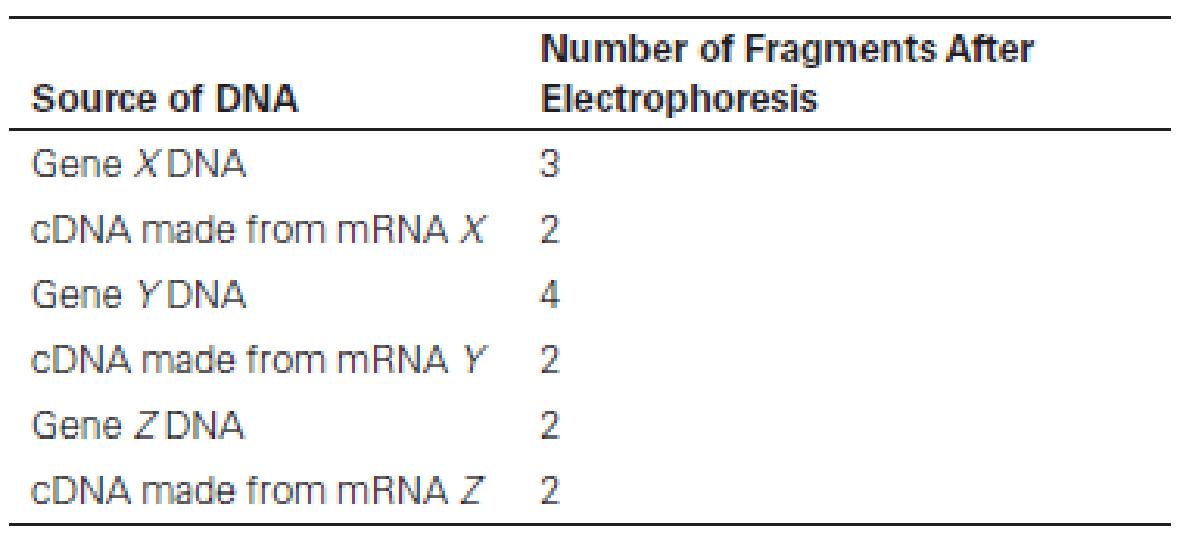
(a) What can you conclude about the number of introns present in gene X?
(b) What can you conclude about the number of introns present in gene Y?
(c) What can you conclude about the number of introns present in gene Z?
Want to see the full answer?
Check out a sample textbook solution
Chapter 21 Solutions
Becker's World of the Cell (9th Edition)
- Tyr- The starting substrate and active site of a Type I topoisomerase is shown below. During this reaction, a small molecule is introduced that removes free hydroxyl groups from DNA (but not protein). Please draw the resulting product under these conditions, including the arrow pushing mechanisms that lead to the product(s). s' CH₂ DNA Base O 111110 H CH₂ Basearrow_forwardPlease answer this asap. Thanks, You have discovered a new plasmid RK21 in a unique bacterial community. As a first step towardunderstanding this plasmid, you digest the plasmid with three restriction enzymes: SspI, XhoI andSmaI. You run the digested plasmid DNA on an agarose gel, along with an uncut sample of theRK21 plasmid DNA as a control.Unfortunately you forget to load a DNA ladder, and obtain the following results. Assumecomplete digestion of all samples or all the digests worked completelyarrow_forwardComplements. The sequence of part of an mRNA is 5'-AUGGGGAACAGCAAGAGUGGGGCCCUGUCCAAGGAG-3' 5'-AUGGGGAACAGCAAGAGUGGGGCCCUGUCCAAGGAG-3' What is the sequence of the DNA coding strand? Of the DNA template strand?arrow_forward
- In the DNA extraction. What is the role of alcohol in the DNA extraction process?arrow_forwardBackward? Bacteriophage T7 helicase moves along DNA in the 5'-to- 3'5'-to-3' direction. Other helicases have been reported to move in the 3'-to-5'3'-to-5' direction. Is there any fundamental reason why you would expect helicases to move in one direction or the other?arrow_forwardHow many sites? A researcher has isolated a restriction endonuclease that cleaves at only one particular 10-base-pair site. Would this enzyme be useful in protecting cells from viral infections, given that a typical viral genome is 50,000 base pairs long? Explainarrow_forward
- Cloning vectors for E Coli. For each of the following items, describe what it is explain the necessity for each in cloning plasmids that are used for the expression (production) of protein. what would happen if this element was omitted? The Multiple Cloning Site (MCS) Origin of replication – (AmpR) antibiotic resistance gene – T7 promoter - T7 termination site- lac I genearrow_forwardPrimer designing: A single-stranded DNA sequence (963 nucleotides) that codes for a hypothetical protein are shown below (lower case shaded blue). 1. Design a pair of forward and reverse primers (~18 nucleotides long each) with EcoRI and BamHI added at 5' and 3' ends, respectively, for the amplification and cloning of this a plasmid with the same restriction sites. gene into GTATCGATAAGCTTGATATCGAATTCatggctaaaggcggagct cccgggttca aagtcgcaat acttggcgct gccggtggcattggccagccccttgcgatgttgatgaagatgaatcctctggtttctgttctacatctatatgatgtagtcaatgcccctggtgtcaccgctgatatta gccacatggacacgggtgctgtggtgcgtggattcttggggcagcagcagctggaggctgcgcttactggcatggatcttattatagtccctgcaggtgttcctcg aaaaccaggaatgacgagggatgatctgttcaaaataaacgcaggaattgtcaagactctgtgtgaagggattgcaaagtgttgtccaagagccattgtcaacctg atcagtaatcctgtgaactccaccgtgcccatcgcagctgaagttttcaagaaggctggaacttatgatccaaagcgacttctgggagttacaatgctcgacgtagt cagagccaatacctttgtggcagaagtattgggtcttgatcctcgggatgttgatgttccagttgttggcggtcatgetggtgtaaccatttgccccttctatctcagg…arrow_forwardRestriction sites of Lambda (A) DNA - In base pairs (bp) The sites at which each of the 3 different enzymes will cut the same strand of lambda DNA are shown in the maps (see figure 3 B-D), each vertical line on the map is where the respective enzymes will cut. A DNA A (bp) 48502 10 000 20 000 30 000 40 000 9162 17 198 B Sal I 7059 14 885 28 338 35 603 42 900 (bp) Hae III 11 826 21 935 29 341 38 016 (bp) 11648 29,624 Eco R1 (bp) 10 592 16 246 28 915 41 864 Figure 3: Restrictrion site map showing the following A) inear DNA that is not cut as reference B) DNA CLt with Sal L C) DNA cut with Hae , D) DNA cut with Eco RI 1. Calculate the size of the resulting fragments as they will occur after digestion and write the sizes on the maps below. Note that linear DNA has a total size of 48 502 bp (see figure 3A). Page 3 of 7 9162 17 198 Sal i (bp) 7059 14 885 28 338 35 603 42 900 Hae I (bp) 11 826 21 935 29 341 38 016 11648 29,624 Eco R1 (bp) 10 592 16 246 28 915 41 864arrow_forward
- transformation and CRISPR. In your own words, briefly describe two differences between these technologies. (For example, these can be differences between their outcomes, procedures, reagents, or something else.)arrow_forwardYes or no? No explanation. rna polymerase are recruiting to start transcription but promoters are DNA sequence. Taq polymerase is enzyme and can synthesize dna at 72 degrees. dna reads 5 to 3 and polymerase reads template 3 to 5.arrow_forwardHi, help please. Which of the following is TRUE regarding RNA editing? a .The coding sequence is altered in the chromosome b. More than one answer choice is correct c. The mRNA is altered by Guide RNAs d. Translation first takes place, following by altering of the coding sequencearrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning

