
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Textbook Question
Chapter 21, Problem 18P
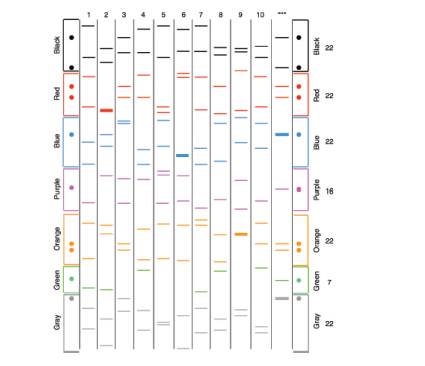
The following figure shows the FBI-style analysis of the genomic DNA of 10 people (1–10), and also of hair found at a crime scene left by the murderer [***]. This analysis involves the PCR amplification of SSR loci, each from a different (nonhomologous) chromosome. The PCR primers are for each SSR locus are labeled with a unique fluorescent molecule. Some bands are thicker because relatively more of the corresponding PCR product was obtained. The figure has dots aligned on both sides to help you find the crucial bands; it will help to use a straight-edge as a guide. The numbers at right are the total number of copies of the SSR locus among the population of 11 samples.
| a. | Are any of individuals 1–10 the murderer? If so, which one? |
| b. | Are any of the loci on the X chromosome? If so, identify this (these) locus (loci) by color. |
| c. | Are any of the loci on the Y chromosome? If so, identify this (these) locus (loci) by color. |
| d. | Are any of individuals 1–10 probable relatives of the murderer? If so, identify this person and describe the degree of relationship to the criminal. |
| e. | One of individuals 1–10 is aneuploid. Write the identity of this person, their sex (M or F), the type of aneuploidy involved, whether the nondisjunction occurred in the mother or father of this person or you can’t tell, and whether the nondisjunction occurred during meiosis I or meiosis II or that you can’t tell. |
| f. | What is the probability that any random male in the United States would share the same genotype as the murderer (the match probability)? Assume that all 11 DNA samples analyzed in the diagram are together representative of the U.S. population as a whole. Show what numbers you would multiply to do this calculation. |
| g. | Explain why the assumption in part (f) that the sample is representative cannot be completely accurate. How might the inaccuracy of this assumption affect the match probability? |
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
The following figure shows the FBI-style analysis of the genomic DNA of 10 people (1-10), and also of hair found at a crime scence left by the murderer [***]. This analysis involves the PCR amplification of SSR loci, each from a different (nonhomologus) chromosome. The PCR primers are for each SSR locus are labeled with a unique fluorescent molecule. Some bands are thicker because relatively more of the corresponding PCR product was obtained. The figure has dots aligned on both sides to help you find the crucial bands; it will help to use a straight-edge as a guide. The numbers at right are the total number of copies of the SSR locus among the population of 11 samples.
e. What is the probability that any random male in USA would share the same genotype as the murderer (the match probability)? Assume that all 11 DNA samples analyzed in the diagram are together representative of the USA population as a whole. Show what numbers you would multiply to do this calculation.
The following figure shows the FBI-style analysis of the genomic DNA of 10 people (1-10), and also of hair found at a crime scence left by the murderer [***]. This analysis involves the PCR amplification of SSR loci, each from a different (nonhomologus) chromosome. The PCR primers are for each SSR locus are labeled with a unique fluorescent molecule. Some bands are thicker because relatively more of the corresponding PCR product was obtained. The figure has dots aligned on both sides to help you find the crucial bands; it will help to use a straight-edge as a guide. The numbers at right are the total number of copies of the SSR locus among the population of 11 samples.
Are any of the loci on the Y chromosome? If so, identify this (these) locus (loci) by color.
The following figure shows the FBI-style analysis of the genomic DNA of 10 people (1-10), and also of hair found at a crime scence left by the murderer [***]. This analysis involves the PCR amplification of SSR loci, each from a different (nonhomologus) chromosome. The PCR primers are for each SSR locus are labeled with a unique fluorescent molecule. Some bands are thicker because relatively more of the corresponding PCR product was obtained. The figure has dots aligned on both sides to help you find the crucial bands; it will help to use a straight-edge as a guide. The numbers at right are the total number of copies of the SSR locus among the population of 11 samples.
Are any of the loci on the X chromosome? If so, identify this (these) locus (loci) by color.
Chapter 21 Solutions
Genetics: From Genes to Genomes
Ch. 21 - Choose the best matching phrase in the right...Ch. 21 - When an allele is dominant, why does it not always...Ch. 21 - A population with an allele frequency p of 0.5 and...Ch. 21 - In a certain population of frogs, 120 are green,...Ch. 21 - Which of the following populations are at...Ch. 21 - A dominant mutation in Drosophila called Delta...Ch. 21 - A large, random mating population is started with...Ch. 21 - Prob. 8PCh. 21 - Alkaptonuria is a recessive autosomal genetic...Ch. 21 - Two hypothetical lizard populations found on...
Ch. 21 - It is the year 1998, and the men and women sailors...Ch. 21 - a. Alleles of genes on the X chromosome can also...Ch. 21 - In 1927, the ophthalmologist George Waaler tested...Ch. 21 - The equation p2 2pq q2> = 1 representing the...Ch. 21 - A gene has two alleles A frequency = p and a...Ch. 21 - Some people can taste the bitter compound...Ch. 21 - Androgenetic alopecia pattern baldness is a...Ch. 21 - The following figure shows the FBI-style analysis...Ch. 21 - Why is the elimination of a fully recessive...Ch. 21 - Tristan da Cunha is a group of small islands in...Ch. 21 - Small population size causes genetic drift because...Ch. 21 - Three basic predictions underlie genetic drift in...Ch. 21 - A mouse mutation with incomplete dominance t =...Ch. 21 - In Drosophila, the vestigial wings recessive...Ch. 21 - In a population of infinite size, three loci A, B,...Ch. 21 - You have identified an autosomal gene that...Ch. 21 - In Europe, the frequency of the CF allele causing...Ch. 21 - An allele of the G6PD gene acts in a recessive...Ch. 21 - Explain why evolutionary biologists monitor...Ch. 21 - Tiny foxes live on the Channel Islands off the...Ch. 21 - What is the most straightforward evidence at the...Ch. 21 - In March 2013, the American Journal of Human...Ch. 21 - If you go back 40 generations into your biological...Ch. 21 - In Fig. 21.17, to what part of the world does...Ch. 21 - Predict the DNA sequences at the four nodes...Ch. 21 - A cladogram not drawn to scale for the taxonomic...Ch. 21 - As noted in Fig. 21.22, humans now living in...Ch. 21 - As of this writing in 2016, no Neanderthal-derived...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- A cloning vector map is shown below. EcoRI Bam Ban Hind P-galactosidase Amp Bam Bam EcoRI Ori C Which restriction site is best for inserting a DNA fragment for selection of chimeric plasmid containing colonies? 1) They're all equally good. 2) Hindll 3) EcoRI 4) BamHIarrow_forwardDNA samples from four individuals were cleaved with the same MW restriction endonuclease. The DNA fragments were separated by gel clectrophoresis, transferred to a membrane, and hybridized with a 12 kb 10 kb DNA probe complementary to a region between sites C and D (see 8 kb hybridization line). The image of the southern blot shows the labeled DNA bands and 6 kb molecular weight (MW) markers. The lane labels I, II, III, and IV -5 kb correspond to individuals I, II, III, and IV. Assume that fragments such as C-D and C-E are clearly resolved in this gel system. Fragment sizes are as given: A-B is 4 kb, B-C is I kb, C-D is 5 kb, and D-E is 650 bp. Individual I has five cleavage sites (A, B, C, D, and E) for the restriction endonuclease. DNA homologous to probe Which individual has at least one point mutation that eliminates restriction site C only? O II IV cannot be determined II Which individual has at least one point mutation that climinates restriction sites B and C? III O IV cannot be…arrow_forwardThe figure below represents the size of various SSRS that are used for forensic analysis. The bars corresponding to each locus represent the range of size of the various alleles of the locus. 100 bp 200 bp 300 bp D8S1179 D21S11 D7S820 CSF1PO D3S1358 TH01 D13S317 D16S539 D2S1338 D199433 VWA TPOX D18S51 FGA D5S818 III Each color in the figure represents a set of molecular markers that can be analyzed simultaneously. Which of the following pairs of markers, which are not grouped together in the figure, could be analyzed simultaneously if they were the only two markers being studied? FGA and CSF1PO 400 bp THO and VWA THO and TPOX O TOPX and FGAarrow_forward
- As the leading scientist in a biomedical science laboratory, it is a requirement to give advice to your lab assistants when they are having problems with their experiments. What advice would you give to your assistants that are having the following problems: After performing a polymerase chain reaction (PCR) and agarose gel electrophoresis to confirm the presence of the C01 gene of 750bp. 2.1. They observe no band appearing on an agarose gel. What would be your conclusion? 2.2. They observe three bands of different sizes that resemble a smear on the gel. Advice 2.3. They observe a single band on the gel and conclude that the PCR product is an exact copy of the original template DNA. Would you support their condusion? Explain. 2.4. Explain how PCR can be used to detect infectious agents in diagnoses of diseases.arrow_forwardA PCR reaction was performed to amplify the CHE2 gene. The PCR primers used were designed to amplify bp 936-4,635 of a 10,908 bp plasmid. Calculate the expected PCR product size (in bp).arrow_forwardPCR primers Below is a 300 base pair fragment of DNA. The top strand is written in the 5' to 3' direction. The bottom strand is written 3' to 5'. There are also two primer sequences; both primers are written 5' to 3'. Note that we are displaying a double-stranded DNA fragment, but primers will only bind to one of the two displayed strands. 5' ACCGȚAGCTATATGCTATCGTGACGTATCGGCGCATTAAȚCGGGATCGAT 3 50 3' TGGCÁTCGATATACOATAGCACTOCATAGCCGCGTAATTÀGCCCTAGCTÀ 5' 5' AGCTÇGCTAGCAGGAGAGAȚATCGÇTCATAGCTCCGATCGATGCCGCTAA 3 3' TCGAGCG ATCGTCCTCTCTÁTAGCGAGTATCGAGÓCTAGCTACGGCGATİ 5' 100 5' TATAGCTCTÇTGCGGATATÇGCATATACCẠ AGGCCCTACGTATGTAGCTA 3 150 3' ATATČGAGAGACOCCTATAGCGTATATGGTTCCGGGATGČATACATCGAŤ 5' 5 TGCGTATATÇGGAGAGTCCTGGATATGGAGCTTGACTGCAGAGAGCTCGA 3 200 3' ACGCÁTATAGCCTCICAGGÁCCTATACCTCGAACTGACGTCTCTCGAGCT 5' 5' TATGCGCTTAGGCCGTATATGCTTGGGGAAAGCTCTATGTATGCTATGTG 3 3. ATACGCGAATCCGGCATATACGAACCCCTÍTCGAGATACATACGATACAC 5' 250 5' TGCATGTGCTATGCAACGTTCOGATTGCGȚAGCAGTAATAGCGCCGATTG 3 300 3'…arrow_forward
- After restriction enzymes cut, they contain unpaired bases. Type II restriction enzymes leave ends that may be 5' overhanging, 3' overhanging, or blunt. In all cases each end is left with a 3' OH and a 5' phosphate. All blunt ends, and any complementary overhanging ends may be re-ligated with T4 DNA ligase, as long as at least one 5'- phosphate is present. In the tables below G^AATTC means that the end after cutting with enzyme will be: -----G 3' -----CTTAA 5' GTGCA^C means that the end will be: -----GTGCA 3' -----C 5' Which RE’s from table below have a 5’ overhang? Which ones have a 3’ Overhang? AccI GT^CGAC BamHI G^GATCC ClaI AT^CGAT NsiI ATGCA^T PstI CTGCA^G BglII A^GATCT TaqI T^CGAarrow_forwardFor the following short sequence of double stranded DNA, design primers (just ~ 3-4 bases) and show 2 copy cycles of PCR (refer to figure 13.25) for the amplification of this sequence of DNA (so that you have 4 double stranded DNA). 5’- GGTATTGGCTACTTACTGGCATCG- 3’ 3’- CCATAACCGATGAATGACCGTAGC- 5’arrow_forwardWhy is it possible to visualize a PCR product on an agarose gel even if the template genome is present at such a low concentration that it cannot be seen?arrow_forward
- Mention all the enzymes and reagents used in the everse Transcriptase Polymerase Chain Reaction (RT-PCR) technique, with their functions.arrow_forwardA PCR reaction was performed to amplify the XULA3 gene, which is bp 882-5,364 on a plasmid that is 11,719 bp. After the PCR, the product was digested with XhoI. There are XhoI sites on the plasmid at bp 1,434, 4,655, and 7,368. Calculate the size(s) that would result when the product is digested with XhoI. Then enter the size of the largest fragment (in bp).arrow_forwardThe short sequence shown below is part of a gene of interest. You decide to amplify this part of the gene by PCR. One primer has been provided by your lab supervisor as shown 3 GAGAGC 51 5'TGAGCAGCGATGCCGATGTTGCCAATGCAGCTCTCG-3'| Which of following additional primers would you need to make? OA S'ACTGCT'3 OB. S'ACGAGT3 OC. SUGAGCA'S D. STGAGCA'3arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
DNA Use In Forensic Science; Author: DeBacco University;https://www.youtube.com/watch?v=2YIG3lUP-74;License: Standard YouTube License, CC-BY
Analysing forensic evidence | The Laboratory; Author: Wellcome Collection;https://www.youtube.com/watch?v=68Y-OamcTJ8;License: Standard YouTube License, CC-BY