
Concept explainers
To analyze:
The steps of biochemical pathway showing the sequence of different compounds and enzymes on the basis of the given data.
Given:
Bacterial strains mutated for five different enzymes (1, 2, 3, 4, and 5) were grown on a variety of media, which was substituted with different compounds involved in the pathway. Table 1 represents the mutant strain growth on the medium having different compounds required in the biochemical pathway.
Table 1: Growth of different mutants on medium containing different compounds of the biochemical pathway.
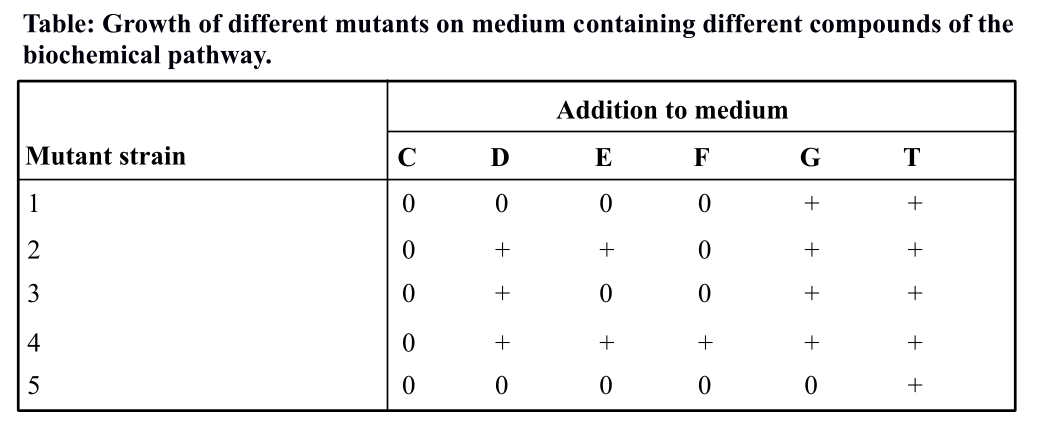
In the table shown above, compound ‘C’ or chorismate is the precursor in the biochemical pathway and ‘T’ or tryptophan is the end product. Compound D, E, F, and G are intermediates. The ‘+’ represents growth when the indicated compound was added to the growth medium, and ‘0’ represents the absence of growth.
Introduction:
In a wild-type bacterium, the synthesis of tryptophan takes place through a series of enzymatic reactions starting from the precursor, chorismate. In this experiment, to deduce the biochemical pathway, the bacterial strains have been mutated for different enzymes and the growth of the bacteria in presence of different compounds has been recorded.
Want to see the full answer?
Check out a sample textbook solution
Chapter 14 Solutions
Life: The Science of Biology
- Consider the following simple regulatory pathways. Assume the full pathway is shown. A- E- B- F- C- G- D- 1 A H- 2 B || L You identify several null mutations (a complete deletion of the gene). For each mutant (indicated with a - sign), determine whether the final product (I, J, K or L) is inducible, uninducible, or constitutive. 3 C 4 D- [Choose ] [Choose ] [Choose ] [Choose ] [Choose ] [Choose ] [Choose ] E [Choose ] F G I H || J Karrow_forwardConsider the following simple regulatory pathways. Assume the full pathway is shown. A- E- B- F- C- G- D- 1 A H- 2 B || L You identify several null mutations (a complete deletion of the gene). For each mutant (indicated with a - sign), determine whether the final product (I, J, K or L) is inducible, uninducible, or constitutive. 3 C 4 D inducible inducible constitutive uninducible constitutive inducible inducible E uninducible F G H > I > J Karrow_forwardIn this problem, just put the order of the intermediates on the pathway, starting with P and ending with Z and explain your reasoning. (So to get you started, notice the class 1 mutants—and there is only mutant in this class, which is mutant #5—won’t grow on minimal medium, but will grow if you give it substance Z. However, giving it substances, p, w, x, or y won’t help. This means that the gene that is mutated lies on the pathway in a place that the Z substances is to its right and all the other substances are to its left. The easiest next one to look at is the line with 2 pluses, and then the line with 3 pluses, and then the line with 4 pluses).arrow_forward
- You generate mutants in the metabolic pathway for starlase. You conduct some complementation tests (after testing for dominance of course) and come up with the following results: 1 2 4 6 1 + + + 2 + 3 + 4 + 5 6 a. How many complementation groups are there? [Select] b. You conduct some additional experiments to elucidate the starlase metabolic pathway. Your results are shown below. Use this information alongside information from the complementation table above to place the intermediates in the correct order on the pathway. (HINT: use the complementation groups from the table above to help you consolidate information on the tables below. Reference practice question 3 from today's lecture for help). Addition to minimal medium Mutant None starlase P 1 + + 2 + + 4 + + 5 + 6 Precursor --> [ Select ] [ Select ] [ Select ] --> starlase c. Mutant 4 has a loss-of-function mutation for which enzyme in the starlase synthesis pathway? [ Select ] E1 E2 ЕЗ Е4 Precursor > Intermediate 1→ Intermediate…arrow_forwardA classic way to isolate thymidylate synthase–negative mutants of bacteriais to treat a growing culture with thymidine and trimethoprim. Most ofthe cells are killed, and the survivors are greatly enriched in thymidylatesynthase–negative mutants.(a) What phenotype would allow you to identify these mutants?(b) What is the biochemical rationale for the selection? (That is, why are themutants not killed under these conditions?)(c) How would the procedure need to be modified to select mammalian cellmutants defective in thymidylate synthase?arrow_forwardCompounds A, B, C, and D are known to be intermediates in the pathway for production of protein E. To determine where the block in protein-E production occurred in each individual, the various intermediates were given to each individual’s cell in culture. After a few weeks of growth with the intermediate, the cells were assayed for the production of protein E. The results for each individual’s cells are given in the following table. A plus sign means that the protein E was produced after the cells were given the intermediate listed at the top of the column. A minus sign means that the cells still could not produce protein E even after being exposed to the intermediate at the top of the column. a) If an individual who is homozygous for the mutation found in individual 2 and heterozygous for the mutation found in individual 4 mates with an individual who is homozygous for the mutation found in individual 4 and heterozygous for the mutation found in individual 2, what could the…arrow_forward
- Clary Leonhart used the pET vector system to express her prokaryotic amylase enzyme. She added peptone into her culture broth of BL21 (DE3) Escherichia coli strain. At the end of the experiment, she discovered that her protein was not expressed. She repeated three more times but her protein of interest was still not produced. (i) (ii) Explain the reason why Clary failed to obtain her protein of interest and suggest a solution to troubleshoot this problem. Clary plans to express her protein along with a polyhistidine-tag, or better known by its trademarked name IIis-tag. Explain the importance of His-tag in protein work.arrow_forwardA classic way to isolate thymidylate synthase-negative mutants of bacteria is to treat a growing culture with thymidine and trimetho- prim. Most of the cells are killed, and the survivors are greatly enriched in thymidylate synthase-negative mutants. (a) What phenotype would allow you to identify these mutants? (b) What is the biochemical rationale for the selection? (That is, why are the mutants not killed under these conditions?) (c) How would the procedure need to be modified to select mam- malian cell mutants defective in thymidylate synthase?arrow_forwardA classic way to isolate thymidylate synthase-negative mutants of bacteria is to treat a growing culture with thymidine and trimethoprim. Most of the cells are killed, and the survivors are greatly enriched in thymidylate synthase-negative mutants. (a) What phenotype would allow you to identify these mutants? (b) What is the biochemical rationale for the selection? (That is, why are the mutants not killed under these conditions?) (c) How would the procedure need to be modified to select mammalian cell mutants defective in thymidylate synthase?arrow_forward
- Consider the following simple regulatory pathways. Assume the full pathway is shown. A- E- B- F- C- G- D- 1 H- A You identify several null mutations (a complete deletion of the gene). For each mutant (ind with a - sign), determine whether the final product (I, J, K or L) is inducible, uninducible, or constitutive. 2 B 3 C 4 D inducible inducible constitutive uninducible constitutive inducible inducible E uninducible F G H > > >arrow_forward. Seven E. coli mutants were isolated. The activity ofthe enzyme β-galactosidase produced by cells containing each mutation alone or in combination with othermutations was measured when the cells were grown inmedium with different carbon sources.Lactose +Glycerol Lactose GlucoseWild type 0 1000 10Mutant 1 0 10 10Mutant 2 0 10 10Mutant 3 0 0 0Mutant 4 0 0 0Mutant 5 1000 1000 10Mutant 6 1000 1000 10Mutant 7 0 1000 10F′ lac from mutant 0 1000 101/ mutant 3F′ lac from mutant 0 10 102/ mutant 3Mutants 3 + 7 0 1000 10Mutants 4 + 7 0 0 0Mutants 5 + 7 0 1000 10Mutants 6 + 7 1000 1000 10Assume that each of the seven mutations is one andonly one of the genetic lesions in the following list.Identify the type of alteration each mutation represents.a. superrepressorb. operator deletionc. nonsense (amber) suppressor tRNA gene (assumethat the suppressor tRNA is 100% efficient in suppressing amber mutations)d. defective CRP–cAMP binding sitee. nonsense (amber) mutation in the β-galactosidase genef.…arrow_forwardIn the: A mutated TBP protein Explain: (a) What is the process affected? (b) What is the Effect on the process? (c) Does it affect prokaryotes, eukaryotes or both?arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





