
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 9, Problem 22P
You use the primer 5′ GCCTCGAATCGGGTACC 3′ to sequence part of the human DNA insert of a recombinant DNA molecule made with a plasmid vector. The result of the automated DNA sequence analysis is shown here. The height of the peaks is unimportant. (A = green; C = purple; G = black; T = red)
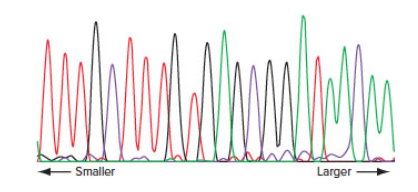
| a. | Write the sequence of all the |
| b. | Is the sequence you wrote in (a) part of the new DNA strand that was synthesized in the sequencing reaction or part of the template strand used in the sequencing reaction? |
| c. | How did you know how to design the primer you would need for the sequencing reaction? Diagram the recombinant DNA molecule to be sequenced, indicating the human and vector sequences, the position and orientation of the primer, and the position and orientation of the new DNA that would be synthesized during the sequencing reaction, using Fig. 9.7 as a guide. |
| d. | Show the full sequence of the smallest DNA molecule that would be synthesized in the sequencing reaction and that would contain dideoxyG (ddG). Indicate the 5′-to-3′ orientation of this molecule and the location of the ddG. |
| e. | How would the data differ from that shown if you accidentally left the dATP out of the reaction? |
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Design a pair of primers to amplify the entire length of the following 45 base pair sequence.Make each primer 14 bases long. Write the sequences of the primers in 5' to 3' order.(Hint: It will help for you to write out BOTH strands of the DNA sequence listed below.5'-GATGCCCGTTGGATAAATTGGGCGTCTAGAATCGGTCACACTTAG-3'
The short DNA shown below is to be sequenced. Using your knowledge of how the Sanger method
works, in the gel diagram, draw in the bands that will appear when DNA polymerase is added to the
reaction along with the four different nucleotide mixtures indicated. Note that some of these mixtures are
not what would normally be used in a sequencing reaction. Dideoxynucleatides (ddNTPs) are added in
relatively small amounts. The asterisk represents a radioactive label.
*5'
- 3'-ОН
3' –
-- ACGACGCAGGACATTAGAC-5'
Nucleotide mixtures:
A. DATP, DTTP, dCTP, DGTP, ddTTP (given)
B. DATP, ATTP, dCTP, AGTP, ddATE
C. dTTP, dGTP. ACTP, ddCTP, ddATP
D. DATP, dCTP, dTTP, ddGTP
A
в с D
| || ||
You were going to sequence a rice DNA fragment whose sequence was only know at one end, as shown below.
5’ AAACGATCGAGTCGCATCCAAAATCGATACCC—unknown region
3’ TTTGCTAGCTCTGCGTAGGTTTTAGCTATGGG—unknown region
After several tries, you obtained a beautiful sequencing image as shown here:
The worked out well partially because you had designed a primer for sequencing the unknown region according to the following guideline:
Tm is 55 – 60°C.
Ensures primer had a appropriate melting temperature for PCR ans sequencing.
The GC content of the primer is the same as the genome/template (rice = 60%, human/Drosophila = 45-50%).
A same nucleotide cannot be more than 2 in a row, e.g. CCC, GGGGG, AAA.
The secondary structure of the primer must be none or weak.
No primer dimers (The primer anneals to itself).
3’ end is the most important: it should not end in A, preferably ends in GG, GC, CG or CC
This website can help you design the primer: http://www.oligoevaluator.com/OligoCalcServlet…
Chapter 9 Solutions
Genetics: From Genes to Genomes
Ch. 9 - Match each of the terms in the left column to the...Ch. 9 - For each of the restriction enzymes listed below:...Ch. 9 - The calculations of the average restriction...Ch. 9 - The DNA molecule whose entire sequence follows is...Ch. 9 - Why do longer DNA molecules move more slowly than...Ch. 9 - Agarose gels with different average pore sizes are...Ch. 9 - The following picture shows the ethidium...Ch. 9 - The linear bacteriophage genomic DNA has at each...Ch. 9 - Consider a partial restriction digestion, in which...Ch. 9 - The text stated that molecular biologists have...
Ch. 9 - a. What is the purpose of molecular cloning? b....Ch. 9 - a. DNA polymerase b. RNA polymerase c. A...Ch. 9 - Is it possible that two different restriction...Ch. 9 - A plasmid vector pBS281 is cleaved by the enzyme...Ch. 9 - A recombinant DNA molecule is constructed using a...Ch. 9 - Suppose you are using a plasmid cloning vector...Ch. 9 - Prob. 17PCh. 9 - The lacZ gene from E. coli encodes the enzyme...Ch. 9 - Your undergraduate research advisor has assigned...Ch. 9 - Which of the enzymes from the following list would...Ch. 9 - You use the primer 5 GCCTCGAATCGGGTACC 3 to...Ch. 9 - a. To make a genomic library useful for sequencing...Ch. 9 - Problem 15 showed part of the sequence of the...Ch. 9 - Eukaryotic genomes are replete with repetitive...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- A facility says they need 15 μL of a 40 ng/μL solution of plasmid DNA for sequencing. The typical yield for a DNA miniprep 5 μg eluted in 50 μL of solution. What do you need to do (for example dilution) to send the appropriate amount to the facility? Show all math work with an explanation.arrow_forwardThe restriction endonuclease NciI recognizes and cuts the five-base-pair sequence 5’- CC(G/C)GG-3’ [where (G/C) means either G or C will work at that position]. (1) How often, on average, would this sequence occur in random DNA? Assume the DNA contains 25% each of A, G, T & C. (2) After digestion, Nci1 leaves a one-base 5’ overhang. Write/draw the cut site/digested products.arrow_forwardThe partial sequence of one strand of a double-stranded DNA molecule is 5'-GACGAAGTGCTGCAGAAAGTCCGCGTTATAGGCATGAATTCCTGAGG -3' EcoRI is a restriction enzyme that cleaves after G in the sequence 5'-GAATTC-3'. PstI is a restriction enzyme that cleaves after A in the sequence 5'-CTGCAG-3'. Write the sequence of both strands of the DNA fragment created when this DNA is cleaved with both EcoRI and PstI. The first strand of your duplex DNA fragment should be derived from the given strand sequence. 5'- -3' 3'- -5'arrow_forward
- You are analyzing the region of DNA shown below to determine how many AATG repeats are present. To do so, you must amplify the entire region of AATG repeats. Design primers of 16 bases each so they anneal outside the region of interest. More than one primer pair is possible, but just give one. 5’-ACTGGCACAGAACAGGCACTTAGGAATGAATGAATGAATGAATGAATGAATGACCTGTGTGGTTCCCAGTTCCTCC-3’ 3’-TGACCGTGTCTTGTCCGTGAATCCTTACTTACTTACTTACTTACTTACTTACTGGACACACCAAGGGTCAAGGAGG-5’arrow_forwardA linear piece of DNA that is 14 kb long is cut first by EcoRI alone, then by SmaI alone, and finally, by both EcoRI and SmaI together. The following results are obtained: Draw a map of the EcoRI and SmaI restriction sites on this 14-kb piece of DNA, indicating the relative positions of the restriction sites and the distances between them.arrow_forwardIn a Sanger DNA sequencing reaction (dideoxy method) you use the primer 5′-GCATATA-3′ to sequence a DNA strand that starts with the sequence 3′-CGTATATCCCTACGTTGG-5′ (consider the strand about 100-nucleotide long). You try your sequencing reaction three times, but every time you make a mistake, as indicated below. What would be the outcome in each case? Justify your answers. (a) You forgot to add dCTP (b) You forgot to add ddCTP (c) Your primer is complementary to two regions in the DNA to be sequenced.arrow_forward
- Examine the DNA sequence shown below. You have been tasked with designing Primers for PCR amplification of the whole fragment shown. Your colleague said that she would design one primer and came up with this sequence – 5’ TTGCATCG 3’. You, being a good scientist, need to confirm that her work is good. Where will this primer bind on the target DNA, and will this primer work as part of a pair to successfully amplify this fragment of DNA? 5’ CGATGCAATCGAGCTATGGCATATCATAAGCGATAGACAGATAGCA 3’ GCTACGTTAGCTCGATACCGTATAGTATTCGCTATCTGTCTATCGT a. It will bind to the bottom strand on the left side of the fragment, and is suitable to amplify the fragment by PCR. b. It will bind to the top strand on the left side of the fragment, but it is unsuitable to amplify the fragment by PCR. c. It will bind to the top strand on the right side of the fragment, but it is unsuitable to amplify the fragment by PCR. d. It will bind to the bottom strand on the right side of the…arrow_forwardBase analysis of a plant DNA shows it to have 15 mole percent adenine (moles per 100 moles total nucleotide). What are the percentages of the other three bases?arrow_forwardFor the following short sequence of double stranded DNA and the given primers, there will be one major duplex DNA product after many cycles (imagine 10 cycles) of PCR. Provide the sequence of this one major duplex product and label the 5’ and 3’ ends of each strand. Sequence to be amplified: 5’- GGTATTGGCTACTTACTGGCATCG- 3’ 3’- CCATAACCGATGAATGACCGTAGC- 5’ Primers: 5’-TGGC-3’ and 5’-TGCC-3’arrow_forward
- A team of scientist is interested in the amplification of a specific DNA fragment in a large plasmid of about 10000 bps. (1) The sequence the scientists are interested in is 5-CATTGATTATTG[ 3-GTAACTAATAAG[ JATCAATTACGGG-3 JTAGTTAATGCCC-5 Where [...] indicates a longer 100bps sequence. Can you suggest two possible primers that the scientists should use to address their need, if they want to be sure they specifically address this region in the entire plasmid? ii) How can the scientist verify that their selection of primers was able to restrict the implementation to only that region of the plasmid?arrow_forwardIn Polymerase Chain Reaction (PCR), the temperature is one of the most important parameters that could influence the efficiency of this technique. Each cycle of this reaction has its own specific temperature. For instance, the denaturation step possesses a temperature of 94 - 98 ℃ to ensure that the double stranded DNA is fully separated. (i) (ii) (iii) Why is the annealing temperature vital in this technique? Explain how will this temperature affects the efficiency of this reaction. Why is Hot Start PCR technique preferred by some researchers? If the primers you purchased possessed the following information. 5'-GGA AAC AGC TAT GAC CAT G-3' Calculate the melting temperature of this primer and estimate the annealing temperature of this primer.arrow_forwardThat's the result of Gel electrophoresis of genomic DNA ( Of genomic DNA extraction experiment), please discuss the results and label and name the image to illustrate the answer? - Marker band sizes in gel: From top (well side) to bottom the bands have the following size in base-pair/bp- 6751,3652,2827,1568,1118,825,630arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Genome Annotation, Sequence Conventions and Reading Frames; Author: Loren Launen;https://www.youtube.com/watch?v=MWvYgGyqVys;License: Standard Youtube License