
Concept explainers
Hydropathy: The Plot Thickens. A hydropathy plot can be used to predict the structure of a membrane protein based on its amino acid sequence and the hydrophobicity values of the amino acids. Hydrophobicity is measured as the standard free energy change, ΔG°′, for the transfer of a given amino acid residue from a hydrophobic solvent into water, in kilojoules per mole (kJ/mol). The hydropathy index is calculated by averaging the hydrophobicity values for a series of short segments of the polypeptide, with each segment displaced one amino acid farther from the N-terminus. The hydropathy index of each successive segment is then plotted as a function of the location of that segment in the amino acid sequence, and the plot is examined for regions of high hydropathy index.
- (a) Why do scientists try to predict the structure of a membrane protein by this indirect means when the technique of X-ray crystallography would reveal the structure directly?
- (b) Given the way it is defined, would you expect the hydrophobicity index of a hydrophobic residue such as valine or isoleucine to be positive or negative? What about a hydrophilic residue such as aspartic acid or arginine?
- (c) Listed below are four amino acids and four hydrophobicity values. Match the hydrophobicity values with the correct amino acids, and explain your reasoning.
Amino acids: alanine, arginine, isoleucine, serine
Hydrophobicity (in kJ/mol): +3.1, +1.0, −1.1, −7.5
- (d) Shown in Figure 7-29 is a hydropathy plot for a specific integral membrane protein. Draw a horizontal bar over each transmembrane segment as identified by the plot. How long is the average transmembrane segment? How well does that value compare with the number you calculated in Problem 7-6c? How many transmembrane segments do you think the protein has? Can you guess which protein this might be?
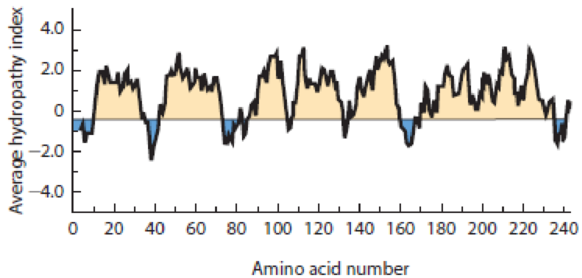
Figure 7-29 Hydropathy Plot for an Integral Membrane Protein. See Problem 7-10d.
QUANTITATIVE That’s About the Size of It. From chemistry, we know that each methylene (—CH2—) group in a straight-chain hydrocarbon advances the chain length by about 0.13 nm. And from studies of protein structure, we know that one turn of an α helix includes 3.6 amino acid residues and extends the long axis of the helix by about 0.56 nm. Use this information to answer the following.
- (a) How long is a single molecule of palmitate (16 carbon atoms) in its fully extended form? What about molecules of laurate (12 C) and arachidate (20 C)?
- (b) How does the thickness of the hydrophobic interior of a typical membrane compare with the length of two palmitate molecules laid end to end? What about two molecules of laurate or arachidate?
- (c) Approximately how many amino acids must a helical transmembrane segment of an integral membrane protein have if the segment is to span the lipid bilayer defined by two palmitate molecules laid end to end?
Want to see the full answer?
Check out a sample textbook solution
Chapter 7 Solutions
Becker's World of the Cell (9th Edition)
- Suggest a reason why amino acids are usually more soluble at pH extremes than they are at neutral pH. (Note: This does not mean that they are insoluble at neutral pH).arrow_forward. A protein gives, under conditions of buffer composition, pH, and tem- perature that are close to physiological conditions, a molecular weight by sedimentation equilibrium measurements of 140,000 g/mol. When the same protein is studied by SDS gel electrophoresis in the absence or presence of the reducing agent B-mercaptoethanol (BME), the pat- terns seen, respectively, in lanes A and B are observed. Lane C contains standards of molecular weight indicated. From these data, describe the native protein, in terms of the kinds of subunits present, the stoi- chiometry of subunits, and the kinds of bonding (covalent, noncova- lent) existing between subunits. / A B - BME + BME STD Serum albumin, 67,000 Ovalbumin, 43,000 Carbonic anhydrase, 30,000 Trypsin inhibitor, 20,000 10 2. 4. Co cm migratedarrow_forwardA spherical cell with the diameter of 10uMhas a protein concentration of 20 mg/ml. Determine the number of protein molecules within the cell if the molecular weight of an average protein is 50,000 daltons (g/mol). Recall that Avogadro's number is NA 6.0221367×1023 molecules/mol. =arrow_forward
- Titration curve of an unknown amino acid The graph below shows a curve, which was obtained following titration of an unknown amino acid. Include a suitable descriptive title stating the identity of the unknown amino acid Use rectangles to precisely outline the regions in the graph where ionisable groups show buffering activity (base/acid ratio 1:10 to base/acid ratio 10:1); clearly associate the name of the ionisable group with the buffering regions; indicate within the graph all observable titration mid-points and all observable titration end-points and indicate estimates of the pKa of the three functional groups (do not provide pka values from the literature, you need to read the pka from the titration curve provided). Note that it is not possible to estimate the pka with more than one decimal precision due to limited resolution of the shown graph.arrow_forwardA large chaperone protein complex GroEL is approximately 16 nm in diameter. When it is dissolved in water at 300 K, estimate the average time it will take for GroEL to diffuse a distance of 500 nm (0.5 micron). The viscosity of water is 10-3 Pa*s.arrow_forwardPredict the number of bands and apparent mol. wt. of the following proteins on SDS gels. 1. A trimeric protein containing three chains, each with a molecular weight of 60,000 Da (60 kDa).arrow_forward
- Under what pH conditions can a protein not bind to the beads in a column? pH = -pKa pH = pI pH = 7 pH = pKa In size exclusion/gel filtration chromatography, the elution order is dependent upon Molecular weight Concentration Overall Charge Enzymatic Activityarrow_forward7. Specificity of membrane transporters. A protein that transports amino acids across the cell membrane was found to bind only a few amino acids efficiently. To find the specificity, many different amino acids and substrate analogs were used as competitive inhibitors in transport studies at pH 5.9 with L-histidine (Km = 10 μM). The Ki values calculated from Lineweaver-Burk plots are shown in the table below. Comparing the structures of L-histidine and the competitive inhibitors (for which most of them, you should know their structures!), what can you conclude about the characteristics of molecules that this transport protein binds at its active site? K; (10-€ M) Amino acid or analog L-Lys L-Arg Gly L-Asp D-His Histamine Dehydrourocanate D-Arg 2 3 285 450 340 390 285 355 HN- + HN + Structure -ΝΗ -NH3 histamine ΝΗ -COO™ dehydrourocanatearrow_forwardThe activity of a pore-forming membrane protein, PorinA1, was tested in different lipid bilayers at 60 °C. Each bilayer is composed of a single type of phosphoglyceride. The four different phosphoglycerides used are shown in the table below together with the corresponding average hydrophobic thickness of the bilayer formed: Phosphoglyceride Bilayer hydrophobic thickness ( A) Dilauroylphosphatidylcholine (DLPC) 30.7 Dimyristoylphosphatidylcholine (DMPC) 34.2 Dipalmitoylphosphatidylcholine (DPPC) 38.1 Distearoylphosphatidylcholine (DSPC) 42.2 PorinA1 is found to have the greatest activity when embedded in DPPC lipid bilayers. Which of the following values (in A) would be the best estimate for the length of the hydrophobic region of PorinA1? O 38 O 51 O 31 O 42 O 38arrow_forward
- A spherical cell with the diameter of 10uMhas a protein concentration of 20 mg/ml. Determine the number of protein molecules within the cell if the molecular weight of an average protein is 50,000 daltons (g/mol). Recall that Avogadro's number is N₁ = 6.0221367×1023 molecules/mol.arrow_forwardaffinity of a protein-protein or protein-ligand interaction can be described by the Dissociation Constant, Kd (written below). Consider a protein P and its inhibitor, I. I inhibits P's activity when bound to it: koff _ [A][B] Dissociation Constant: Ka = koN [AB] Question When [I] is 10-7 M, 99% of P's activity is inhibited. What is the Kd of this Protein- Inhibitor interaction?arrow_forwardTrue or false: The contributions to the change in Gibbs free energy of binding for a lipid ligand in the lipid bilayer to a membrane protein are going to be similar to those driving a water soluble ligand binding a free floating protein. Briefly explain your reasoning.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





