
Concept explainers
Problem Set
Inside or Outside? From Figure 7-22, we know that exposed regions of membrane proteins can be labeled with 125I by the lactoperoxidase (LP) reaction. Similarly, carbohydrate side chains of membrane glycoproteins can be labeled with 3H by oxidation of galactose groups with galactose oxidase (GO) followed by reduction with tritiated borohydride (3H—BH4). Noting that both LP and GO are too large to penetrate into the interior of an intact cell, explain each of the following observations made with intact erythrocytes.
(a) When intact cells are incubated with LP in the presence of 125I and the membrane proteins are then extracted and analyzed on SDS–polyacrylamide gels, several of the bands on the gel are found to be radioactive.
(b) When intact cells are incubated with GO and then reduced with 3H—BH4, several of the bands on the gel are found to be radioactive.
(c) All of the proteins of the plasma membrane that are known to contain carbohydrates are labeled by the GO/3H—BH4 method.
(d) None of the proteins of the erythrocyte plasma membrane that are known to be devoid of carbohydrate is labeled by the LP/125I method.
(e) If the erythrocytes are ruptured before the labeling procedure, the LP procedure labels virtually all of the major membrane proteins.
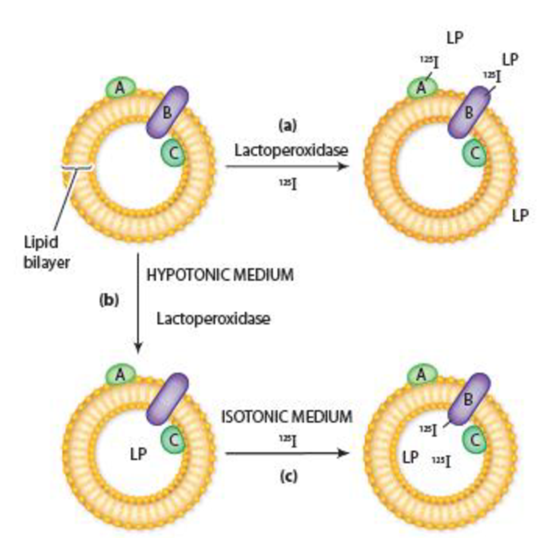
Figure 7-22 A Method for Labeling Proteins Exposed on One or Both Surfaces of a Membrane Vesicle. (a) In the presence of 125I, lactoperoxidase (LP) labels membrane proteins exposed on the outer surface of membrane vesicles (i.e., proteins A and B). If membrane vesicles are (b) first incubated in a hypotonic medium to make them permeable to LP and (c) then transferred to an isotonic solution containing 125I but no external LP, the membrane proteins exposed on the inner membrane surface (i.e., proteins B and C) become labeled.
Want to see the full answer?
Check out a sample textbook solution
Chapter 7 Solutions
Becker's World of the Cell (9th Edition)
- Activity (% Max.) Using your knowledge of aspartyl proteases, develop a plausible explanation for why protein activity changes when pH is increased or decreased from pH 4, as shown in the graph. 06 08 09 10+ 4. 9. Hd Which statements are true? The protein is active when one asp residue is protonated and the other is deprotonated. O Both asp residues are protonated at more basic pH values. O Active aspartyl proteases have only one asp residue in the active site. Only one asp side chain is protonated at low pH (below pH 3.5). Both asp side chains are deprotonated at higher pH values.arrow_forwardP3D.2 In biological cells, the energy released by the oxidation of foods is stored in adenosine triphosphate (ATP or ATP“).The essence of ATP's action is its ability to lose its terminal phosphate group by hydrolysis and to form adenosine diphosphate (ADP or ADP): ATP* (aq) + H,O() → ADP* (aq) + HPO (aq) + H,O*(aq) At pH = 7.0 and 37°C (310K, blood temperature) the enthalpy and Gibbs energy of hydrolysis are A,H =-20kJ mol and A,G=-31 kJ mol", respectively. Under these conditions, the hydrolysis of 1 mol ATP“(aq) results in the extraction of up to 31kJ of energy that can be used to do non- expansion work, such as the synthesis of proteins from amino acids, muscular contraction, and the activation of neuronal circuits in our brains. (a) Calculate and account for the sign of the entropy of hydrolysis of ATP at pH = 7.0 and 310K. (b) Suppose that the radius of a typical biological cell is 10µm and that inside it 1x 10ʻ ATP molecules are hydrolysed each second. What is the power density of…arrow_forwardGlycerophospholipids Phosphatidylethanolamine 3. In case the cell is in a state requiring large amount of ATP to support energy-requiring reactions/pathways, assuming that you have 1 mole of each of the said lipids are catabolized and complete oxidized, will the total net ATP yield from these two lipids be higher or lower than the sum of the net ATPs generated from each fatty acid components? Justify your answer in biochemical terms and using 5 sentences or less.arrow_forward
- Glycolipids: What are the similarities and differences in the biosynthesis of glycosphingolipids, GPI anchors and N-linked glycans?arrow_forwardDrawing the Structure of a Glycopeptide (Integrates with Chapters 4and 5.) Consider the peptide DGNILSR, where N has a covalentlylinked galactose and S has a covalently linked glucose. Draw thestructure of this glycopeptide, and also draw titration curves for theglycopeptide and for the free peptide that would result from hydrolysis of the two sugar residues.arrow_forward. Separating Glycated Hb From Normal Hb (Integrates with Chapters 5and 6.) Human hemoglobin can react with sugars in the blood(usually glucose) to form covalent adducts. The a-amino groups ofN-terminal valine in the Hb b-subunits react with the C-1 (aldehyde)carbons of monosaccharides to form aldimine adducts, whichrearrange to form very stable ketoamine products. Quantitation ofthis “glycated hemoglobin” is important clinically, especially fordiabetic individuals. Suggest at least three methods by which glycatedHb (also referred to as HbA1c) could be separated from normal Hband quantitatedarrow_forward
- 1. pH Effects a. In the enzyme mechanism of lysozyme, two acidic amino acid residues, Asp52 and Glu35, are critical for catalytic activity. If we assume normal side chain pKa values for Asp (pKar = 3.90) and for Glu (pKar = 4.07), what proportion of enzyme molecules will have both Asp52 and Glu35 in the correct ionization state at pH 5.0 (the pH optimum for lysozyme)? b. Are the traditional pKa values likely to be correct within the protein? What pKa changes might be present within lysozyme?arrow_forwardDe novo purine biosynthesis is a 10-step pathway that builds the purine rings one atom, or one group of atoms, at a time. The pathway converts 5-phosphoribosyl-1-pyrophosphate (PRPP) to inosine monophosphate (IMP). Identify the nine intermediates on the pathway diagram. 5-phosphoribosyl-1-amine PRPP glycinamide ribonucleotide formylglycinamide rimuclocticke N-succinyl-5-aminoimidazole-4- carboxamide ribonucleotide 5-aminoimidazole-4- carboxamide ribonucleotide Incorrect -CH, H H OH Gn Glu -CH. H 0. CH H glycine + ATP HN HC HC ADP+P OH OH OH 0 -PFI HN (P-Ribose Formy-THF HN THE H HN -Ribose -ATP 6-Ⓡ CH₂ HN HC CH, P-Ribose -ATP ADP+P NH, ADP+P HN NH -Ribose NH, N-formylaminoimidazole-4- carboxamide ribonucleotide formylglycinamidine ribonuclcotide S-aminoimidazole ribonuclotidie ATP ADP+Pi HCO, carboxyanincimidazole ribonucleotide -Ribose H -Ribose H₂O Formyl-THF- ASP Fumarate HC THE 0 N. "NH₂ Ⓒ-Abse NH, NH, -ATP NH -N-CH COO ADP+PI Coo IMP COO NH, 000 Answer Bankarrow_forwardHi - I have the following as a question in my biochemistry course and I am just lost on where to start. Thanks! Cyt cb562 will form a tetramer in the presence of Zn+2 or in the absence of Zn+2, it will form a trimer. For both the tetramer and the trimer, suggest the identity of three different amino acids that could be on the interface. Explain how your selections facilitate these interactions.arrow_forward
- Need help The β chains of HbA and HbS were treated with trypsin, and the sequence of the N-terminal tryptic peptides are as given below. Do these peptides separate from each other in an electric field if the pH is 7.0? Explain in detail the reasoning behind your answer and include your calculations for the charge of each peptide in your answer. HbA: Val-His-Leu-Thr-Pro-Glu-Glu-Lys HbS: Val-His-Leu-Thrarrow_forwardProtein,nfolded Proteinţolded Under certain conditions, a specific protein has the following thermodynamic parameters for going from the unfolded state to the folded state: AH = – 300 kJ/mol, and TAS = – 400 kJ/mol. Under those conditions, the equilibrium would lie more toward neither would predominate - the fraction of total protein that's folded would = fraction unfolded. the folded state the unfolded state O It is impossible to predict the position of the equilibrium from AH and TAS.arrow_forward. The process of a protein folding from an inactive unfolded structure to the active folded structure can be represented by the following equation: unfolded protein = folded protein The values of AH and AS° for the folding of the protein lysozyme are: AH = -280 kJ/mol AS = -790 J/mol · K (a) Calculate the value of AG for the folding of lysozyme at 25 °C. (b) At what temperature would you expect the unfolding of lysozyme to become favorable?arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

