
Concept explainers
The phage P1 is used as a generalized transducing phage in an experiment combining a donor strain of E. coli of genotype leu+phe+ala+ and a recipient strain that is leu-phe-ala-. In separate experiments, transductants are selected for leu+ (experiment A), for phe+ (experiment B), and for ala+ (experiment C). Following selection, transductant genotypes for the unselected markers are identified.
a. What compound or compounds are added to the minimal medium to select for transductants in experiments A, B, and C?
b. Selection experiment results below show the frequency of each genotype.
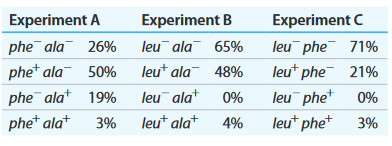
c. Determine the order of genes on the donor chromosome.
d. Diagram the crossover events that form each of the transductants in experiment A.
e. In experiment B, why are there no transductants with the genotype leu- ala+?
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Austin Taylor and Edward Adelberg isolated some new strains of Hfr cells that they then used to map several genes in Escherichia coli by using interrupted conjugation. In one experiment, the researchers mixed cells of Hfr strain AB‑312, which were xyl+ mtl+ mal+ met+ and sensitive to phage T6, with F− strain AB‑531, which was xyl− mtl− mal− met− and resistant to phage T6. The cells were allowed to undergo conjugation. At regular intervals, the researchers removed a sample of cells and interrupted conjugation by killing the Hfr cells with phage T6. The F− cells, which were resistant to phage T6, survived and were then tested for the presence of genes transferred from the Hfr strain. The results of this experiment are shown in the graph. On the basis of these data, give the order of the xyl, mtl, mal, and met genes on the bacterial chromosome and the minimum distances between them in minutes. The origin of transfer is represented by the red triangle. The distances between genes are not…arrow_forwardE. coli cells are simultaneously infected with two strains of phage λ. One strain has a mutant host range, is temperature sensitive, and produces clear plaques (genotype h st c); another strain carries the wildtype alleles (genotype h+ st+ c+). Progeny phages are collected from the lysed cells and are plated on bacteria. The following numbers of different progeny phages are obtained: Progeny phage genotype Number of plaques h+ c+ st+ 321 h c st 338 h+ c st 26 h c+ st+ 30 h+ c st+ 106 h c+ st 110 h+ c+ st 5 h c st+ 6 a. Determine the order of the three genes on the phage chromosome. b. Determine the map distances between the genes. c. Determine the coefficient of coincidence and the interferencearrow_forwardIn recombination studies of the rII locus in phage T4, what is the significance of the value determined by calculating phage growth in the K12 versus the B strains of E. coli following simultaneous infection in E. coli B? Which value is always greater?arrow_forward
- Austin Taylor and Edward Adelberg isolated some new strains of Hfr cells that they then used to map several 200 mal+ genes in Escherichia coli by using interrupted conjugation. 150 In one experiment, the researchers mixed cells of Hfr strain AB-312, which were xyl* mtl* mal* met* and sensitive to phage T6, with F strain AB-531, which was xyl mtl mal met and resistant to phage T6. The 100 mt/+ cells were allowed to undergo conjugation. At regular intervals, the researchers removed a sample of cells and 50 met+ interrupted conjugation by killing the Hfr cells with phage T6. The F cells, which were resistant to phage T6, survived and were then tested for the presence of 0. 20 40 60 80 100 genes transferred from the Hfr strain. The results of this experiment are shown in the graph. Time of sampling (minutes) On the basis of these data, give the order of the xyl, mtl, mal, and met genes on the bacterial chromosome and the minimum distances between them in minutes. The origin of transfer is…arrow_forwarda. Bacteriophage P22 was used in generalized transduction experiments to infect the Salmonella typhimurium donor strains described in the table below. The resulting phage lysates were then used to infect the S. typhimurium recipient strains listed in the table. In each cross, a phenotype was selected for one of the three genetic markers studied (str, aceA, thrA), and then replicates were performed to select the corresponding recombinants for the other two markers. The results are given in the following table: Recipient strain Selected phenotype Selected recombinants Donor strain str thrA aceAt thrA str aceA+ strs thrA+ aceA thrA+ str aceA Str Ace+ Str ThrA ThrA+ ThrA ThrA+ Ace Ace+ str: gene involved in streptomycin resistance, aceA: gene involved in the use of acetate as a carbon source, thrA: gene involved in the biosynthesis of threonine. Number 60 40 95 5 10 90 What are the selective media used in these three transduction experiments, on the one hand to obtain the selected…arrow_forwardIn E. coli, the gene bioD+ encodes an enzyme involved in biotin synthesis, and galK+ encodes an enzyme involved in galactose utilization. An E. coli strain that contained wild-type versions of both genes was infected with P1 phage, and then a P1 lysate was obtained. This lysate was used totransduce (infect) a strain that was bioD− and galK−. The cellswere plated on a medium containing galactose as the sole carbonsource for growth to select for transduction of the galK+ gene.This medium also was supplemented with biotin. The resultingcolonies were then restreaked on a medium that lacked biotin tosee if the bioD+ gene had been cotransduced. The following resultswere obtained:What topic in genetics does this question address?arrow_forward
- In E. coli, the gene bioD+ encodes an enzyme involved in biotin synthesis, and galK+ encodes an enzyme involved in galactose utilization. An E. coli strain that contained wild-type versions of both genes was infected with P1 phage, and then a P1 lysate was obtained. This lysate was used totransduce (infect) a strain that was bioD− and galK−. The cellswere plated on a medium containing galactose as the sole carbonsource for growth to select for transduction of the galK+ gene.This medium also was supplemented with biotin. The resultingcolonies were then restreaked on a medium that lacked biotin tosee if the bioD+ gene had been cotransduced. The following resultswere obtained:What information do you know based onthe question and your understanding of the topic?arrow_forwardShown below are the complementation test results involving 4 independently isolated lethal mutants in a bacteriophage. Complementation was assayed by simultaneouly infecting bacteria with two phage strains, each with a different mutation, neither of which could alone lyse the cells. In the table below, a "+" indicates the strains complemented each other and therefore lysed open the bacteria. A "0" indicates no complementation and therefore no cell lysis occurred. Test pair Results 1___2___3___4 1,2 + 1 0 + + 0 1,3 + 2 0 + + 1,4 0 3 0 + 2,3 + 4 0 2,4 + 3,4 + How many genes are there? a. 3 b.1 c. 2 d. 4arrow_forwardShown below are the complementation test results involving 4 independently isolated lethal mutants in a bacteriophage. Complementation was assayed by simultaneouly infecting bacteria with two phage strains, each with a different mutation, neither of which could alone lyse the cells. In the table below, a "+" indicates the strains complemented each other and therefore lysed open the bacteria. A "0" indicates no complementation and therefore no cell lysis occurred. Test pair Results 1___2___3___4 1,2 + 1 0 + + 0 1,3 + 2 0 + + 1,4 0 3 0 + 2,3 + 4 0 2,4 + 3,4 + Which mutants are in the same gene? . a. 2, & 3 b.1, 2, 3 & 4 c.1 & 4 d.1, 2 & 4arrow_forward
- A microbial geneticist isolates a new mutation in E. coliand wishes to map its chromosomal location. She usesinterrupted-mating experiments with Hfr strains andgeneralized-transduction experiments with phage P1.Explain why each technique, by itself, is insufficient foraccurate mapping.arrow_forwardTwo mutations that affect plaque morphology in phages (a− and b −) have been isolated. Phages carrying both mutations (a− b−) are mixed with wild-type phages (a+ b+) and added to a culture of bacterial cells. Once the phages have infected and lysed the bacteria, samples of the phage lysate are collected and cultured on plated bacteria. The following numbers of plaques are observed: Plaque phenotype Number a+ b+ 2043 a+ b− 320 a− b+ 357 a− b− 2134 What is the frequency of recombination between the a and b genes?arrow_forwardA bacterium of genotype a+ b+ c+ d+ is the donor in a cotransformation mapping. The recipient is a-b-c-d-. Data from the transformed cells is shown below. What is the order of the genes? a+ and b+ 5 a+ and c+ 2 a+ and d+ 0 b+ and c+ 0 b+ and d+ 0 c+ and d+ 5 cbad cadb bcda acbd bacdarrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





