
Campbell Essential Biology (7th Edition)
7th Edition
ISBN: 9780134765037
Author: Eric J. Simon, Jean L. Dickey, Jane B. Reece
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 12, Problem 15PS
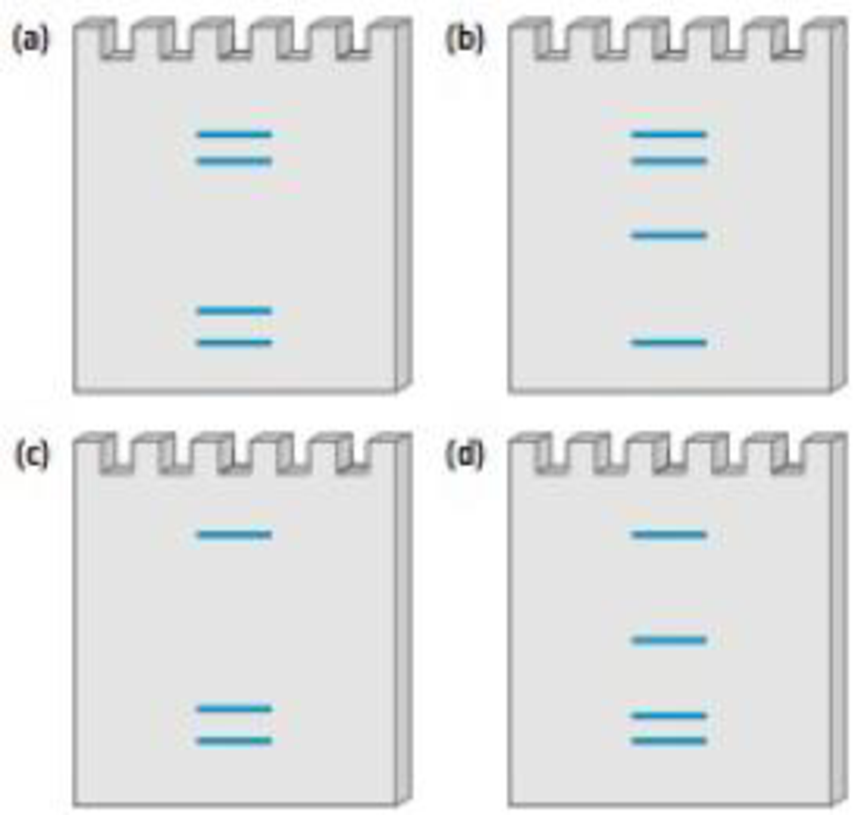
Listed below are 4 of the 13 genome sites used to create a standard DNA profile. Each site consists of a number of short tandem repeats: sets of 4
| Chromosome number | Genetic site | # of repeats |
| 3 | D3S13S8 | 4 |
| 5 | DSS818 | 10 |
| 7 | D7S820 | 5 |
| 8 | D8S1179 | 22 |
Imagine that you perform a PCR procedure to create a DNA profile for this individual. Which of the following four gels correctly represents the DNA profile of this person?
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Listed below are 4 of the 13 genome sites used to create a standard DNA profile. Each site consists of a number of short tandem repeats: sets of 4 nucleotides repeated in a row within the genome. For each site, the number of repeats found at that site for this individual are listed: Imagine you perform a PCR procedure to create a DNA profile for this individual. Which of the following four gels correctly represents the DNA profile of this person?
Design a pair of primers to amplify the entire length of the following 45 base pair sequence.Make each primer 14 bases long. Write the sequences of the primers in 5' to 3' order.(Hint: It will help for you to write out BOTH strands of the DNA sequence listed below.5'-GATGCCCGTTGGATAAATTGGGCGTCTAGAATCGGTCACACTTAG-3'
For the following short sequence of double stranded DNA and the given primers, there will be one major duplex DNA product after many cycles (imagine 10 cycles) of PCR. Provide the sequence of this one major duplex product and label the 5’ and 3’ ends of each strand.
Sequence to be amplified:
5’- GGTATTGGCTACTTACTGGCATCG- 3’
3’- CCATAACCGATGAATGACCGTAGC- 5’
Primers: 5’-TGGC-3’ and 5’-TGCC-3’
Chapter 12 Solutions
Campbell Essential Biology (7th Edition)
Ch. 12 - Suppose you wish to create a large batch of the...Ch. 12 - A carrier that moves DNA from one cell to another,...Ch. 12 - Prob. 3SQCh. 12 - A paleontologist has recovered a bit of organic...Ch. 12 - Why do DNA fragments containing STR sites from...Ch. 12 - What feature of a DNA fragment causes it to move...Ch. 12 - After a gel electrophoresis procedure is run, the...Ch. 12 - Name the steps of the whole-genome shotgun method.Ch. 12 - Put the following steps of human gene therapy in...Ch. 12 - For each statement, identify which major theme is...
Ch. 12 - Prob. 11IMTCh. 12 - For each statement, identify which major theme is...Ch. 12 - Some scientists once joked that when the DNA...Ch. 12 - Interpreting Data When comparing genomes from...Ch. 12 - Listed below are 4 of the 13 genome sites used to...Ch. 12 - In the not-too-distant future, gene therapy may be...Ch. 12 - Today, it is fairly easy to make transgenic plants...Ch. 12 - 18. In October 2002, the government of the African...Ch. 12 - From 1977 to 2000, 12 convicts were executed in...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Below are 9 possible primer pairs.● Determine which primer pair is the best choice.● Explain why the other primers are not good choices.● Calculate the Tm for each primer. Underline or highlight the region of DNA for the primer pair you chose as the best.Forward 1: 5’ gaaataattttgtttaactttaag 3’ Tm =Reverse 1: 5’ gtttaagacaaaatagtctgg 3’ Tm =Forward 2: 5’ gtaactcagctttcaggtcg 3’ Tm =Reverse 2: 5’ tctcggaatgttgcaacagc 3’ Tm =Forward 3: 5’ agattagcggatcctacctg 3’ Tm =Reverse 3: 5’ atgtgtaatcccagcagcag 3’ Tm =Forward 4: 5’ cattgattatttgcacggcg 3’ Tm =Reverse 4: 5’ aaaatcttctctcatccgcc 3’ Tm =Forward 5: 5’ tccataagattagcggatcc 3’ Tm =Reverse 5: 5’ tgcaagcttggctgttttgg 3’ Tm =Forward 6: 5’ gatcctacctgacgcttttta 3’ Tm=Reverse 6: 5’ aaataatgaattcgagctcggt 3’ Tm =Forward 7: 5’ataaaaaaatcgagataaccgtt 3’ Tm =Reverse 7: 5’aggtcgactctagaggatc 3’ Tm =Forward 8: 5’ctacctgttccatggccaac 3’ Tm=Reverse 8: 5’ ttcgggcatggcactcttg 3’ Tm=Forward 9: 5’ tccataagattagcggatcc 3’ Tm =Reverse 9: 5’…arrow_forwardIn addition to the standard base-paired helical structures, DNA can form X-shaped hairpin structures called cruciforms in which most bases are involved in Watson–Crick pairs. Such structures tend to occur at sequences with inverted repeats. Draw the cruciform structure formed by the DNA sequence TCAAGTCCACGGTGGACTTGC.arrow_forwardThe short DNA shown below is to be sequenced. Using your knowledge of how the Sanger method works, in the gel diagram, draw in the bands that will appear when DNA polymerase is added to the reaction along with the four different nucleotide mixtures indicated. Note that some of these mixtures are not what would normally be used in a sequencing reaction. Dideoxynucleatides (ddNTPs) are added in relatively small amounts. The asterisk represents a radioactive label. *5' - 3'-ОН 3' – -- ACGACGCAGGACATTAGAC-5' Nucleotide mixtures: A. DATP, DTTP, dCTP, DGTP, ddTTP (given) B. DATP, ATTP, dCTP, AGTP, ddATE C. dTTP, dGTP. ACTP, ddCTP, ddATP D. DATP, dCTP, dTTP, ddGTP A в с D | || ||arrow_forward
- A Sanger product of the sequencing of a template DNA is presented +ddTTP +ddATP +ddCTP +ddGTP Mononucleotide Pentanucleotide Trinucleotide Dinucleotide 11-nucleotide Hexanucleotide Octanucleotide Tetranucleotide 16-nucleotide Decanucleotide Nonanucleotide Heptanucleotide 17-nucleotide 15-nucleotide 13-nucleotide 12-nucleotide 18-nucleotide 20-nucleotide 14-nucleotide 19-nucleotide 21-nucleotide 1. Determine the sequence of template DNA 2. Determine the mRNA sequence from this template DNA 3. Determine the the sequence of protein product derived from that DNAarrow_forwardFigure 9-22 shows the first steps in the process of making a DNA microarray, or DNA chip, using photolithography. Describe the remaining steps needed to obtain the desired sequences (a different fournucleotide sequence on each of the four spots) shown in the first panel of the figure. After each step, give the resulting nucleotide sequence attached at each spot.arrow_forwardAfter restriction enzymes cut, they contain unpaired bases. Type II restriction enzymes leave ends that may be 5' overhanging, 3' overhanging, or blunt. In all cases each end is left with a 3' OH and a 5' phosphate. All blunt ends, and any complementary overhanging ends may be re-ligated with T4 DNA ligase, as long as at least one 5'- phosphate is present. In the tables below G^AATTC means that the end after cutting with enzyme will be: -----G 3' -----CTTAA 5' GTGCA^C means that the end will be: -----GTGCA 3' -----C 5' Which RE’s from table below have a 5’ overhang? Which ones have a 3’ Overhang? AccI GT^CGAC BamHI G^GATCC ClaI AT^CGAT NsiI ATGCA^T PstI CTGCA^G BglII A^GATCT TaqI T^CGAarrow_forward
- A 13-nucleotide section of an autoradiogram from a Sanger sequencing experiment is depicted in the image below. Based on the band pattern observed here, write out the sequence of both the complementary strand generated during the experiment, and the template strand that is being analyzed. Be sure to clearly indicate the 5' and 3' ends of each. ddATP ddGTP ddCTP || | | ddTTP | 3' 5' Tyne a short answer in the space provided belowarrow_forwardA geneticist is interested in determining the locations of methylated cytosines within a fragment of DNA. She treats some copies of the fragment with sodium bisulfite and leaves some copies untreated. She then sequences the treated and untreated copies of the fragment and obtains the following results. Give the original sequence of the DNA fragment and indicate the locations of methylated cytosines. Sequence without treatment: — AATTGCCCGATCGATTAAGCCA — Sequence with treatment: — AATTGTTTGATCGATTAAGCTA —arrow_forwardThe following is the base sequence of DNA that codes for first eight amino acids of the β chain of hemoglobin. The β chain of hemoglobin contains a total of 147 amino acids so this is a small part of the entire gene. DNA Template Strand: TACCACGTGGACTGAGGACTCCTC 1. What is the minimum number of DNA nucleotides in this whole gene? 2. What is the sequence of bases on the strand of DNA that is complementary to the template strand? 3. What mRNA will be formed from the template strand of DNA?arrow_forward
- Align two sequences: horizontal – GGAATGG, vertical – ATG, m=1, mm = 0, g=-1. Use the table below for the NW matrix. Write down and score all optimal global alignments. Complete the NW matrix below and show the alignment paths. Use the arrows and circles for the matrix and paths. Click the shapes and move them using your keyboard arrow keys (or drag). Click the shapes and then right-click to copy/paste or click and use Ctrl + C to copy, then Ctrl + V to paste (in Windows MS Word). 0000 Align and score 4 optimal alignments here. the first line, v sequence in the second line and Each alignment should have h sequence individual scores for each alignment position in the third line.arrow_forwardThat's the result of Gel electrophoresis of genomic DNA ( Of genomic DNA extraction experiment), please discuss the results and label and name the image to illustrate the answer? - Marker band sizes in gel: From top (well side) to bottom the bands have the following size in base-pair/bp- 6751,3652,2827,1568,1118,825,630arrow_forwardGiven the DNA sequence of the restriction enzyme: gi|6329444|dbj|AB034757.1| Hynobius retardatus mRNA for larval beta-globin, complete cds GCAGAATCTGACTCAAGAAATCCCTCCTCACCCAACACCACCAGCAGCCATGGTTCACTGGACAGCAGAGGAGAAGGCAGCCATCAGCTCTGTGTGGAAGCAGGTGAACGTGGAGAGCGATGGACAGGAGGCCCTGGCCAGGTTGCTGATCGTCTACCCCTGGACCCAGAGATACTTCAGCTCTTTTGGGGACCTGTCGAGCCCAGCTGCCATTTGTGCCAACGCCAAGGTCCGTGCCCATGGCAAGAAGGTCCTGTCCGCCCTGGGAGCCGGCGCCAACCACCTGGATGACATCAAAGGCAACTTTGCTGATCTGAGCAAGCTTCACGCAGACACACTCCATGTGGACCCCAATAACTTCCTGCTCCTGGCAAACTGCCTGGTGATCGTCTTGGCCCGCAAGCTGGGAGCCGCCTTCAACCCTCAAGTCCATGCGGCCTGGGAGAAGTTCCTGGCCGTCTCCACCGCGGCTCTGTCCAGAAACTACCACTAGAGACTGGTCTTTGGGTTTAATTCTGTGAACGTCCCTGAGACAAATGATCTTTCAATGTGTAAACCTGTCATTACATCAATAAAGAGACATCTAACAAAAAAAAAAAAAAAAAAAAAAAAAA Identify two blunt-end cutters Identify two sticky-end cutters. For each, Provide the sequence of the Restriction enzyme, Highlight using a specific color where the DNA sequence where the restriction enzyme will cut the DNA Indicate the…arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
QCE Biology: Introduction to Gene Expression; Author: Atomi;https://www.youtube.com/watch?v=a7hydUtCIJk;License: Standard YouTube License, CC-BY