
Concept explainers
(a)
Interpretation:
The smallest unit of the antibody having the specificity and binding affinity of the whole protein should be determined.
Concept introduction:
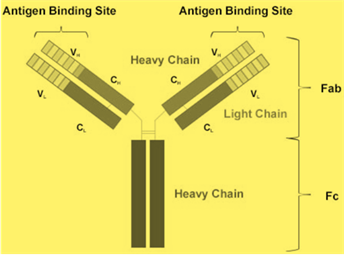
The antibodies are the immunoglobulins (Ig) made up of two types of chains that is a light chain and heavy chain. The immunoglobulin is a class of proteins found in the plasma of the animals. The typical structure of the antibody is given below:
(b)
Interpretation:
A compact single chain that is likely to bind with antigen should be designed.
Concept introduction:
The antibodies are the immunoglobulins (Ig) made up of two types of chains that is a light chain and heavy chain. The immunoglobulin is a class of proteins found inthe plasma of the animals. The antigens are the substance that can elicit an immune response by binding to the antibody.
Want to see the full answer?
Check out a sample textbook solution
- PROTEINS1. What will happen to free tyrosine after being subjected to xanthoproteic analysis (is it positive or negative, include color)2. What will happen to lysine after being subjected to millon's analysis (is it positive or negative, include color)arrow_forward. What physiological effect would you predict from a mutation that replaced with serine the cysteine in the constant part of the immunoglobulin light chain that is involved in disulfide-bond for- mation with the heavy chain? ,arrow_forwardBIOMOLECULES - MULTIPLE CHOICE - Please answer properly QUESTION : The antibody-antigen binding site is different from an active site of an enzyme because an enzyme’s active site. A. contains amino acids without sidechains B. contains modified amino acids C. is complementary to a specific ligand D. catalyzes a chemical reactionarrow_forward
- Describe how the GPCR and MAPK pathways 'cross-talk' or interact with each other. Give support for your statements. Is cross-talk relevant for drug design? Why or why not?arrow_forwardIn SDS-PAGE. What chemical is used to ensure that all protein molecules are coated with a negative charge? IPTG. β-mercaptoethanol. X-Gal. SDS.arrow_forwardExperiment This experiment was designed to analyze the structure of a component (C3c protein) of the complement system, which is involved in the immune response against microorganisms. Purified C3c protein (molecular mass: 145 kd) was incubated in the presence of various concentrations of the reducing agent dithiothreitol (DTT) and then subjected to electrophoresis in a sodium dodecyl sulfate (SDS)-polyacrylamide gel. SDS disrupts noncovalent bonds and polypeptides are separated by size during electrophoresis. The gel was stained with Coomassie Brilliant Blue, a protein dye. The figure shows the molecular masses of intermediates and products generated by DTT treatment. Results: Molecular mass (kd) 145- 102- 75- 43- 27- [DTT] (MM) 10.00 1 2 3 4 5 6 Samples Questions 1. How many polypeptides are present in C3c? Determine their molecular masses from the figure. What was the purpose of using DTT in this experiment? What kind of bonds hold the polypeptides 2. 3. together? 4. How is the 102…arrow_forward
- Please asap. thankyou. The following diagram represents the results of hemoglobin samples of 5 patients (1-5) that were run on an electrophoresis gel at pH 9.2. The first 3 lanes show the control samples, for comparison. Using only the letters "AA" "AS" and "SS", fill in the blanks below indicating the most likely genotype of each patient, based on these protein electrophoresis results. The following diagram represents the results of hemoglobin samples of 5 patients (1-5) that were run on an electrophoresis gel at pH 9.2. The first 3 lanes show the control samples, for comparison. Using only the letters "AA" "AS" and "SS", fill in the blanks below indicating the most likely genotype of each patient, based on these protein electrophoresis results. Patient 1 Patient 2 Patient 3 Patient 4 Patient 5arrow_forwardExperiment This experiment was designed to analyze the structure of a component (C3c protein) of the complement system, which is involved in the immune response against microorganisms. Purified C3c protein (molecular mass: 145 kd) was incubated in the presence of various concentrations of the reducing agent dithiothreitol (DTT) and then subjected to electrophoresis in a sodium dodecyl sulfate (SDS)-polyacrylamide gel. SDS disrupts noncovalent bonds and polypeptides are separated by size during electrophoresis. The gel was stained with Coomassie Brilliant Blue, a protein dye. The figure shows the molecular masses of intermediates and products generated by DTT treatment. Results: Molecular mass (kd) 145- 102- 75- 43- 27- [DTT] (MM) 10.00 1 2 3 4 5 6 Samples Questions 4. How is the 102 kd polypeptide related to the other polypeptide species?arrow_forwardExperiment This experiment was designed to analyze the structure of a component (C3c protein) of the complement system, which is involved in the immune response against microorganisms. Purified C3c protein (molecular mass: 145 kd) was incubated in the presence of various concentrations of the reducing agent dithiothreitol (DTT) and then subjected to electrophoresis in a sodium dodecyl sulfate (SDS)-polyacrylamide gel. SDS disrupts noncovalent bonds and polypeptides are separated by size during electrophoresis. The gel was stained with Coomassie Brilliant Blue, a protein dye. The figure shows the molecular masses of intermediates and products generated by DTT treatment. Results: Molecular mass (kd) 145- 102- 75- 43- 27- [DTT] (MM) 1 2 3 4 5 6 Samples Questions 3. What kind of bonds hold the polypeptides together? |arrow_forward
- Monoclonal antibodies can be conjugated to an insoluble support by chemical methods. Explain how these antibody-bound beads can be exploited for protein purification.arrow_forwardThe F ab fragment of an antibody molecule has essentially the same affinity for a monovalent hapten as does intact IgG. (a) What is the smallest unit of an antibody that can retain the specificity and binding affinity of the whole protein? (b) Design a compact single-chain protein that is likely to specifically bind antigen with high affinity.arrow_forwardproteins. Which of the following will tell you whether a protein would be found in the lumen of the ER? A. You run a hydropathy plot an look for hydrophobic peaks that span 20-30 amino acids B. You isolate microsomes and see whether the proteins are inserted into the membrane of the microsome C. You run a hydropathy plot an look for a lack of hydrophobic peaks that span 20-30 amino acids O D. You do in vitro translation of each protein in the presence or absence of microsomes and look to see whether there is a size change in the presence of microsomes.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
