
Concept explainers
Comparisons of Neanderthal mitochondrial DNA with that of modern humans indicate that they are not related to modern humans and did not contribute to our mitochondrial heritage. However, because Neanderthals and modern humans are separated by at least 25,000 years, this does not rule out some forms of interbreeding causing the modern European gene pool to be derived from both Neanderthals and early humans (called Cro-Magnons). To resolve this question, Caramelli et al. (2003. Proc. Natl Acad Sci. [USA] 100: 6593–6597) analyzed mitochondrial DNA sequences from 25,000–year-old Cro-Magnon remains and compared them to four Neanderthal specimens and a large dataset derived from modern humans. The results are shown in the graph.
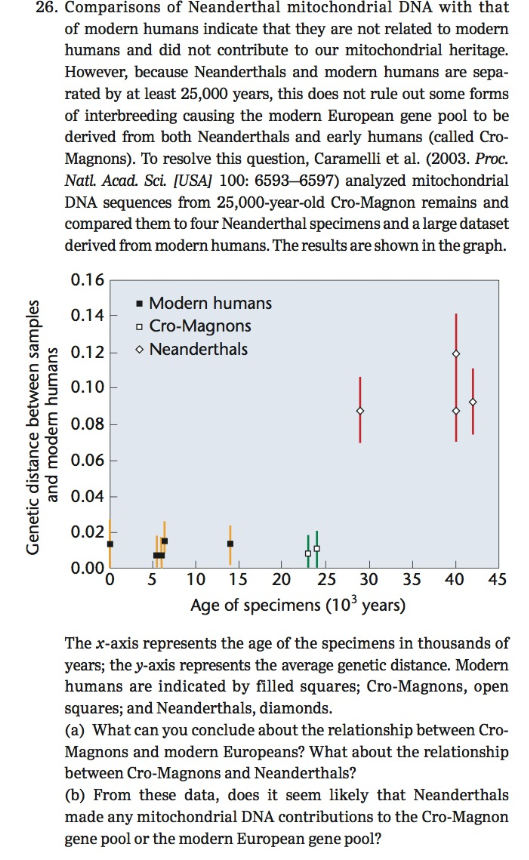
The x-axis represents the age of the specimens in thousands of years; the y-axis represents the average genetic distance. Modern humans are indicated by filled squares; Cro-Magnons, open squares; and Neanderthals, diamonds.
(a) What can you conclude about the relationship between Cro-Magnons and modern Europeans? What about the relationship between Cro-Magnons and Neanderthals?
(b) From these data, does it seem likely that Neanderthals made any mitochondrial DNA contributions to the Cro-Magnon gene pool or the modern European gene pool?
Want to see the full answer?
Check out a sample textbook solution
Chapter 22 Solutions
Essentials of Genetics (9th Edition) - Standalone book
- A number of comparisons of nucleotide sequences among hominidsand rodents indicate that inbreeding may have occurredmore often in hominid than in rodent ancestry. Bakewell et al.(2007. Proc. Nat. Acad. Sci. [USA] 104: 7489-7494) suggest thatan ancient population bottleneck that left approximately 10,000 humans might have caused early humans to have a greaterchance of genetic disease. Why would a population bottleneckinfluence the frequency of genetic disease?arrow_forwardHow, specifically, is the concept of ALLOMETRY relevant to the phylogenetic position of Homo floresiensis? Because if allometry explains the anatomy of Homo floresiensis then we can conclude that it is not separate species but instead a member of our species. Although most mammals on islands go through a process of getting smaller, Homo floresiensis evolved from a smaller ancestor to be bigger, meaning that allometry is an important factor. Mutations in the allometry allele are associated with many of the characteristics of Homo floresiensis. Because Homo floresiensis is so much smaller than other members of the genus Homo, it is important to determine how shape changes associated with smaller size impacted the species. Because Homo floresiensis had both small- and large-bodied forms, variation within the species is in large part dictated by allometry.arrow_forwardAbout 3 million years ago, the Isthmus of Panama (a narrow strip of land connecting North and South America) formed, dividing marine organisms into Pacific and Caribbean populations. Researchers have examined species of snapping shrimp on both sides of the isthmus. Based on the morphological species concept, there appeared to be seven pairs of species, with one species of each pair in the Pacific and the other in the Caribbean. The different species pairs live at somewhat different depths in the ocean. Using mitochondrial DNA sequences, the researchers estimated phylogenies and found that each of these species pairs, separated by the isthmus were indeed each other's closest relatives. The researchers investigated mating in the lab and found that many species pairs were not very interested in courting with each other, and any that did mate almost never produced fertile offspring. Which process led to the formation of the species pairs of Pacific and Caribbean snapping shrimp? sympatric…arrow_forward
- About 3 million years ago, the Isthmus of Panama (a narrow strip of land connecting North and South America) formed, dividing marine organisms into Pacific and Caribbean populations. Researchers have examined species of snapping shrimp on both sides of the isthmus. Based on the morphological species concept, there appeared to be seven pairs of species, with one species of each pair in the Pacific and the other in the Caribbean. The different species pairs live at somewhat different depths in the ocean. Using mitochondrial DNA sequences, the researchers estimated phylogenies and found that each of these species pairs, separated by the isthmus were indeed each other's closest relatives. The researchers investigated mating in the lab and found that many species pairs were not very interested in courting with each other, and any that did mate almost never produced fertile offspring. Use the species concepts to explain how the sister populations on opposite sides of the isthmus are true…arrow_forwardMichael Bunce and his colleagues in England, Canada, and theUnited States extracted and sequenced mitochondrial DNA from fossilsof Haast’s eagle, a gigantic eagle that was driven to extinction 700 yearsago when humans first arrived in New Zealand (M. Bunce et al. 2005.PLOS Biology 3:44–46). Using mitochondrial DNA sequences fromliving eagle species and those from Haast’s-eagle fossils, they createdthe accompanying phylogenetic tree. On this phylogenetic tree, identify(a) all nodes; (b) one example of a branch; and (c) the outgroup.arrow_forwardA group of researchers are trying to determine the relationship between various groups of organisms. The researchers analyze the amino acid sequence of the protein cytochrome c in various groups of organisms. Once they sequence cytochrome c in all organisms, they determine the number of amino acid substitutions in each group. The results can be seen in the graph below, which plots the data with respect to the time since the divergence of the members of paired groups from a common ancestor. A B mammals and reptiles с Number of Amino Acid Substitutions (per 100 amino acids) in Cytochrome, c D 60- 50+ 40+ 30- 20+ 10+ 0 0 O birds and reptiles A graph showing when various groups of organisms diverged from a common ancestor Based on the graph above, can you conclude which of the following organisms are most distantly related? 200 400 600 800 1,000 Time Since Divergence from a Common Ancestor (millions of years) fish and land vertebrates mammals and reptiles birds and mammals insects and…arrow_forward
- How does the fact that all ethnic groups except Africans contain some Neanderthal DNA (1–4 percent of their DNA) support the out-of-Africa hypothesis for the origin of modern humans (Homo sapiens)?arrow_forwardThe phylogenetic tree for vertebrates depicted below was constructed from sequence data for two rRNA mitochondrial genes (12S and 16S). How do the results of this analysis compare with the phylogenetic trees in Figures 32.10 and 32.24? Identify the major clades of vertebrates on the tree depicted below. Source: R. Zardoya and A. Meyer. 1998. Complete mitochondrial genome suggests diapsid affinities of turtles. Proceedings of the National Academy of Sciences, USA 95:1422614231. Copyright 1998 National Academy of Sciences, U.S.A.arrow_forwardThe figure shows a phylogenetic tree of various members of the order Proboscidea, which includes modern elephants. Which of the following claims is best supported by the information in the figure ? a.The mastodon and the Stegodon diverged from their common ancestor 22 million years ago. b.The common ancestor of the African elephant and the mastodon is the Palaeomastodon. c.The mammoth diverged from its most recent common ancestor with African elephants before the mastodon diverged from its most recent common ancestor with Stegodons. d.The Asian and African elephants are the most closely related species shown on the tree.arrow_forward
- Crickets have colonized each island in the Hawaiian Archipelago. Geological data indicates that Kauai is the oldest island in the chain and Hawaii is the youngest. Researchers hypothesized that crickets sequentially colonized islands as they rose out of the ocean and created a cladogram based on molecular relationships to test this idea. 1) Mark & label (as “MRCA”) the most recent common ancestor for all crickets on the island of Kauai. 2) Molokai is roughly equally distant from Oahu as Maui, but not equally related. Are Molokai crickets more closely related to those on Oahu or Maui?_________arrow_forwardReferring to the phylogenetic tree shown above, answer the following questions: 1. How many OTUs are included in the phylogenetic analysis? 2. How many clades are there? 3. What is an autapomorphic trait of the domestic cat? Explain why? 4. What is the shared derived trait (synapomorphy) in the Family Felidae? Explain why?arrow_forwardImagine that you have the DNA sequences from the intron of a gene in three species called A, B, and C. Species A and B are most closely related, while C is more distantly related. The sequences of A and B differ by 18 base pairs, A and C differ by 26 base pairs, and B and C differ by 28 base pairs. Fossils show that species A and B diverged about 1.2 Mya, but there is no fossil evidence as to when the most recent common ancestor of all three species lived. (Draw a simple tree to help you think about the problem) Use the genetic data to estimate that date (most recent common ancestor). HINT = use Eqn 7.1, several times- first to estimate mutation rate. Then to estimate the unknown time since divergencearrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

