
Prescott's Microbiology
11th Edition
ISBN: 9781260211887
Author: WILLEY, Sandman, Wood
Publisher: McGraw Hill
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 17.1, Problem 2MI
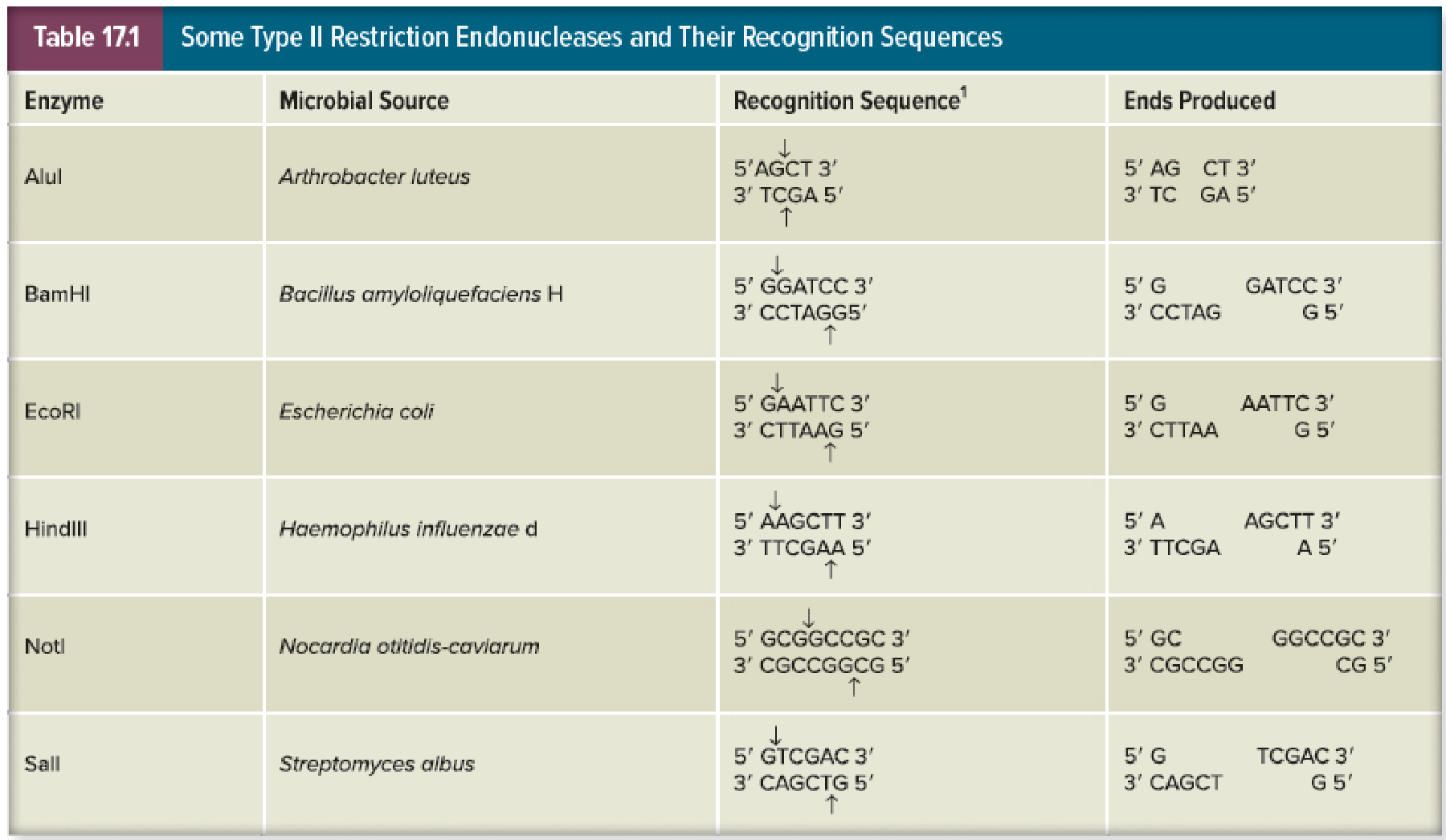
Which of the above enzymes yield blunt ends? Which yield sticky ends with a 5′ overhang? What about a 3′ overhang?
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
What is the principle behind the very specific sequence in donning and doffing PPEs? Explain
What is the reduction division? Why is it necessary??
Draw the Haworth projections of the two anomers of L-gulose. Clearly indicate which anomer is represented.
Chapter 17 Solutions
Prescott's Microbiology
Ch. 17.1 - Examine the uncut piece of DNA shown in the upper...Ch. 17.1 - Which of the above enzymes yield blunt ends? Which...Ch. 17.1 - Prob. 3MICh. 17.1 - What would you conclude if you obtained only blue...Ch. 17.1 - Why must introns be removed from eukaryotic DNA...Ch. 17.1 - Which plasmid is a shuttle vector? Why?Ch. 17.1 - In what ways does the BAC shown here differ from...Ch. 17.1 - Describe restriction enzymes, sticky ends, and...Ch. 17.1 - What is cDNA? Why is it necessary to generate cDNA...Ch. 17.1 - Prob. 3CC
Ch. 17.1 - Prob. 4CCCh. 17.1 - Prob. 5CCCh. 17.2 - Why, after three cycles, are the vast majority of...Ch. 17.2 - Briefly describe the polymerase chain reaction....Ch. 17.2 - Why is PCR used to detect infectious agents that...Ch. 17.2 - How would you use PCR to measure the concentration...Ch. 17.2 - Why is it possible to visualize a PCR product on...Ch. 17.2 - Prob. 5CCCh. 17.3 - Why are long fragments (e.g., 20,000 bp) of...Ch. 17.4 - What special considerations are necessary if one...Ch. 17.4 - Prob. 1CCCh. 17.4 - Prob. 2CCCh. 17.4 - Prob. 3CCCh. 17.4 - You are studying chemotaxis proteins in a newly...Ch. 17.5 - Prob. 1MICh. 17.5 - Prob. 1CCCh. 17.5 - Prob. 2CCCh. 17 - Which of the DNA molecules shown are recombinant?Ch. 17 - Prob. 1RCCh. 17 - Prob. 2RCCh. 17 - Prob. 3RCCh. 17 - Prob. 4RCCh. 17 - Prob. 5RCCh. 17 - Prob. 6RCCh. 17 - Prob. 1ALCh. 17 - Prob. 2ALCh. 17 - Suppose you transformed a plasmid vector carrying...Ch. 17 - You are interested in the activity and regulation...Ch. 17 - Prob. 5ALCh. 17 - Prob. 6ALCh. 17 - Prob. 7AL
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- You generate mutants in the metabolic pathway for starlase. You conduct some complementation tests (after testing for dominance of course) and come up with the following results: 1 2 3 4 5 6 1 2 + 3 + + a. How many complementation groups are there? [Select] 4 + 5 + + + + 6 +arrow_forwardWhich one of these is correct ? And why are the rest incorrect?arrow_forwardYou generate mutants in the metabolic pathway for starlase. You conduct some complementation tests (after testing for dominance of course) and come up with the following results: 1 2 3 4 5 6 1 I 2 + 3 + 4 + + LO 5 a. How many complementation groups are there? [Select] 6 + + + + +arrow_forward
- Given the following complementation chart for holes in Monstera, give me the biochemical (phenotype) pathway. A В D E Holes Class 1 + + + - - Class 2 + + Class 3 + + + Class 4 Class 5 + + - - + +arrow_forwardA classic way to isolate thymidylate synthase–negative mutants of bacteriais to treat a growing culture with thymidine and trimethoprim. Most ofthe cells are killed, and the survivors are greatly enriched in thymidylatesynthase–negative mutants.(a) What phenotype would allow you to identify these mutants?(b) What is the biochemical rationale for the selection? (That is, why are themutants not killed under these conditions?)(c) How would the procedure need to be modified to select mammalian cellmutants defective in thymidylate synthase?arrow_forwardWhich of the follwing enzymes adds incoming deoxyribonucleotid triphosphates to the 3' OH of the growing es/deoxyribonucleoside daughter strand in the 5'-3' direction?arrow_forward
- For E. coli strains with the lac genotypes show below, use a plus sign (+) to indicate the synthesis of β-galactosidase and permease and a minus sign (–) to indicate no synthesis of the proteins.arrow_forwardWhat is a protease enzyme? Why would we treat the strawberry extract with a protease in an extraction where the DNA needed to be subsequently analyzed?arrow_forwardThe intermediates A, B, C, D, E, and F all occur in the same biochemical pathway G is the product of the pathway, and mutations 1 through 7 are all G –, meaning that they cannot produce substance G. The following table shows which intermediates will promote growth in each of the mutants. Arrange the intermediates in order of their occurrence in the pathway at which each mutant strain is blocked. A “+” in the table indicates that the strain will grow if given that substance, an “o” means lack of growth.arrow_forward
- You generate mutants in the metabolic pathway for starlase. You conduct some complementation tests (after testing for dominance of course) and come up with the following results: 1 2 4 6 1 + + + 2 + 3 + 4 + 5 6 a. How many complementation groups are there? [Select] b. You conduct some additional experiments to elucidate the starlase metabolic pathway. Your results are shown below. Use this information alongside information from the complementation table above to place the intermediates in the correct order on the pathway. (HINT: use the complementation groups from the table above to help you consolidate information on the tables below. Reference practice question 3 from today's lecture for help). Addition to minimal medium Mutant None starlase P 1 + + 2 + + 4 + + 5 + 6 Precursor --> [ Select ] [ Select ] [ Select ] --> starlase c. Mutant 4 has a loss-of-function mutation for which enzyme in the starlase synthesis pathway? [ Select ] E1 E2 ЕЗ Е4 Precursor > Intermediate 1→ Intermediate…arrow_forwardWhat different between the generated amplification product and the primer sequences in the CDS sequence?arrow_forwardThe pathway for arginine biosynthesis in Neurospora crassa involves several enzymes that produce a series of intermediates as shown. O O O O ornithine citrulline ARG-E arginosuccinate arginine N-acetylornithine arginine You did a cross between ARG-E ARG-H* and ARG-E* ARG-H¯¯ Neurospora strains and identified an Arg- strain from an NPD tetrad. (Assume that Neurospora forms tetrads in the same way yeast do.) Which compound would rescue growth of this Arg- spore? N-aceltylornithine ARG-F ornithine citrulline ARG-G ARG-H → argininosuccinatearrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning
TISSUE REPAIR Part 1: Repair - Regeneration; Author: ilovepathology;https://www.youtube.com/watch?v=t-5EjlS6qjk;License: Standard YouTube License, CC-BY