
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 17, Problem 5P
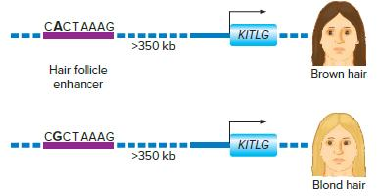
As shown in the following diagram, a single
Expert Solution & Answer
Trending nowThis is a popular solution!

Students have asked these similar questions
The following diagram show what is required for an active promoter of a gene of interest, where:A1 = Activator 1A2 = Activator 2Med = MediatorRep = Repressor
Based on the following data, predict:
Chromatin conformation
Methylation state of the proximal promoter
If protein A1 is present or absent
If protein A2 is present or absent
If the mediator is present or absent
If the repressor is absence or present
If this gene is likely to be transcribed or notPlease note that this is an "all or nothing" bonus question, and no partial credit will be awarded. Selecting all answers will result in zero points awarded. Selecting at least one incorrect answer will result in zero points being awarded.
Question 28 options:
Euchromatin
Heterochromatin
Methylated promoter
Unmethylated promoter
Activator A1 present
Activator A1 absent
Activator A2 present…
Shown below is the genomic structure of the human B-globin
gene. The numbers within the boxes indicate the length in
nucleotides of each region.
Question 6: How many amino acids are present in the wild-type
human B-globin protein?
= exons
= introns
Transcription termination
site (also poly A site)
Promoter
Start of transcription
3'
5'
ATG
50
TẠC
TAA
126
132
|ATT
90
130
222
850
3
5'
Start codon
Stop codon
А. 438
В. 146
C. 620
D. 206
© 2013 John Wiley & Sons, Inc. All rights
reserved.
ennar
region of gene X, which determines the length of the tail in mice, is mutated so
that transcription factors bind it at a much higher affinity compared to the wild-type sequence.
What is the most likely phenotypic outcome?
Tail length will not change because the enhancer is a non-coding sequence
Tail length will increased due to increased activity of the gene's promoter
Tail length will decreased because any mutation will cause a loss-of-function of these regulatory regions
Not just the tail will be enlarged because increased activity of the enhancer will impact many genes
Chapter 17 Solutions
Genetics: From Genes to Genomes
Ch. 17 - For each of the terms in the left column, choose...Ch. 17 - For each of the following types of gene...Ch. 17 - List five events other than transcription...Ch. 17 - Which eukaryotic RNA polymerase RNA pol I, pol II,...Ch. 17 - As shown in the following diagram, a single...Ch. 17 - You have synthesized an enhancerless GFP reporter...Ch. 17 - Prob. 7PCh. 17 - Prob. 8PCh. 17 - A single UAS regulates the expression of three...Ch. 17 - MyoD is a transcriptional activator that turns on...
Ch. 17 - a. Assume that two transcription factors are...Ch. 17 - Prob. 12PCh. 17 - In Problem 12, you identified a genomic region...Ch. 17 - Prob. 14PCh. 17 - Prob. 15PCh. 17 - Genes in both prokaryotes and eukaryotes are...Ch. 17 - Prob. 17PCh. 17 - Lysine 4 of histone H3 H3K4 is methylated in the...Ch. 17 - J.T. Lis and collaborators have developed an...Ch. 17 - Hydatiform moles are growths of undifferentiated...Ch. 17 - Prader-Willi syndrome is caused by a mutation in...Ch. 17 - The human IGF2 gene is autosomal and maternally...Ch. 17 - Follow the expression of a paternally imprinted...Ch. 17 - Reciprocal crosses were performed using two inbred...Ch. 17 - Interestingly, imprinting can be tissue-specific....Ch. 17 - Prob. 26PCh. 17 - A method for detecting methylated CpGs involves...Ch. 17 - Honeybees Apis mellifera provide a striking...Ch. 17 - Consider the experiment in Fig. 17.24, where the...Ch. 17 - A protein or RNA that regulates gene expression in...Ch. 17 - a. How can a single eukaryotic gene give rise to...Ch. 17 - A hunchback gene, a gene necessary for proper...Ch. 17 - You know that the mRNA and protein produced by a...Ch. 17 - You are studying a transgenic mouse strain that...Ch. 17 - Prob. 35PCh. 17 - Scientists have exploited the siRNA pathway to...Ch. 17 - Persimmons Diospyros lotus are dioecious plants,...Ch. 17 - Drosophila females homozygous for loss-of-function...Ch. 17 - The text has discussed the RNA-Seq technique,...Ch. 17 - Researchers know that Fru-M controls male sexual...Ch. 17 - The Drosophila gene Sex lethal Sxl is deserving of...Ch. 17 - Figure 17.29 shows that the Sxl protein binds to...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Consider the two-line model of a gene below: A ΟΥ Y If the promoter is found to the left of the transcription initiation site, where would you find the transcribed region? A +1arrow_forwardDiscuss the following argument: “if the expression of every gene depends on a set of transcription regulators, then the expression of these regulators must also depend on the expression of other regulators, and their expression must depend on the expression of still other regulators, and so on. cells would therefore need an infinite number of genes, most of which would code for transcription regulators.” how does the cell get by without having to achieve the impossible?arrow_forwardA gene encodes a protein with the following amino acid sequence: Met-Lys-Ser-Pro-Ala-Thr-Pro A nonsense mutation caused by a single-base-pair substitution occurs in this gene, resulting in a protein with the amino acid sequence Met-Lys. An intergenic suppressor mutation allows the gene to produce the fulllength protein. With the original mutation and the intergenic suppressor present, the gene now produces a protein with the following amino acid sequence: Met-Lys-Cys-Pro-Ala-Thr-Pro Give the location and nature of the original mutation and of the intergenic suppressor.arrow_forward
- The level of transcription of a gene is tested by creating deletions in the gene as shown in the illustration. These modified genes are tested for their level of transcription: (++) normal transcription levels • (+) low transcription levels (+++) high transcription levels. Which deletion is in an enhancer involved in regulating the gene? Normal gene 2 3 4 ++ Deletion 1 3 4 +++ Deletion 2 1 3 4 ++ Deletion 3 1 3 + Deletion 4 2 4 ++ 4+arrow_forwardYou have been analyzing wing development mutants in Drosophila and have collected the genetic/Western data below. Your epistasis analysis (not shown), suggested your current model that DW-2 is upstream of WL-1. What is a plausible mechanism for how DW-2 regulates WL-1? mutant name wingless-1 doublewing-2 Prated with Ants-Wingless MW standards || | | mutanttype ODW-2 inhibits transcription of WL-1 O DW-2 polyubiquitinates WL-1 null null phenotype Model Ac O DW-2 phosphorylates WL-1 and prevents it from entering the nucleus O DW-2 cleaves WL-1 proteolytically Wings Mode of regulation DW-2 Wings Wingsarrow_forwardDescribe the 4 types of suppressor high copy suppression, bypass suppression, nonsense suppression, protein interaction . in your Own words. Often geneticists look for suppressors to find interactive proteins. Which of the type(s) of suppressors you put for part a will help to identify interacting proteins, and which type(s) will not? What are two (or one, if we don’t get a chance to talk about two of them in class) other techniques (not necessarily “genetic” techniques, but at least, lab techniques) that help to identify identifying proteins?arrow_forward
- A strain of Arabidopsis thaliana possesses a mutation in the APETALA2 gene. As a result of this mutation, much of the 3′ UTR of the mRNA transcribed from the gene is deleted. What is the most likely effect of this mutation on the expression of the APETALA2 gene?arrow_forwardConsider the Rho-dependent terminator sequence 5’CCCAGCCCGCCUAAUGAGCGGCCUUUUUUUU-3’. What affect would a point mutation at any one of the bolded and underlined nucleotides disrupt termination of transcription? Group of answer choices Mutation in one of these nucleotides would disrupt base pairing, preventing the formation of the hairpin and disrupting termination. Mutation in one of these nucleotides would have no affect on base pairing, so the termination hairpin is formed and termination proceeds. Mutation in one of these nucleotides would not disrupt base pairing, but would prevent the formation of the hairpin and disrupt termination. Mutation in one of these nucleotides would disrupt base pairing, but not affect the formation of the hairpin and termination proceeds.arrow_forwardCD3 is a signaling protein that is typically found only in the plasma membrane of immune system T lymphocytes. CD3 is composed of several different polypeptides, including a gamma chain, CD3γ. Scientists analyzed the promoter of the CD3γ chain gene for regulatory sequences that might have positive or negative effects on expression of the gene. The scientists cloned fragments of the CD3γ gene that included the first transcribed nucleotides plus up to 789 nucleotides of upstream regulatory sequences into plasmids in which the gene for the firefly enzyme luciferase immediately follows the fragments. The plasmids were then introduced into a line of T lymphocytes (Figure 1), and the cells were allowed to grow for a short while. Because the regulatory sequences of the CD3γ gene immediately precede the luciferase gene in the plasmids, the activity, either positive or negative, of the regulatory sequences affected the amount of luciferase gene expression by the T lymphocytes. Luciferase…arrow_forward
- RNAP can access the lacz promoter irrespective of whether CAP-CAMP is binding the CAP O True O Falsearrow_forwardIn the bacteriophage T7 system used to express recombinant proteins, the gene of interest is fused to T7 promoter and T7 RNA polymerase is separately cloned into the same cell. What is the main reason this system uses T7 RNA polymerase instead of relying on the bacterial RNA polymerase? To restrict the expression of bacterial protein expression To enhance the amount of recombinant protein expression To enhance the expression of bacterial protein expression To restrict the amount of recombinant protein expression To enable the expression of T7 viral protein expressionarrow_forwardRefer to figure, which shows the distribution of histone H2A.Z on nucleosomes near a transcription start site. What experimental technique would have been used to generate the data for this figure? Briefly describe the operation of this technique.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning

Biology (MindTap Course List)
Biology
ISBN:9781337392938
Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. Berg
Publisher:Cengage Learning
QCE Biology: Introduction to Gene Expression; Author: Atomi;https://www.youtube.com/watch?v=a7hydUtCIJk;License: Standard YouTube License, CC-BY