
Concept explainers
The electrophoresis gel shown in part (a) is from a DNase footprint analysis of an operon transcription control region. DNA sequence analysis of a
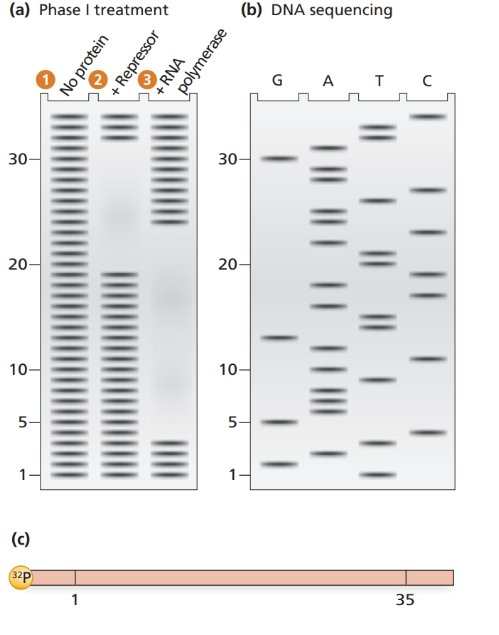
a. Determine the DNA sequence of the region examined.
b. Locate the regions of the sequence protected by repressor protein and by RNA polymerase.
Want to see the full answer?
Check out a sample textbook solution
Chapter 12 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- The following shows the genotype of a partial diploid bacterial cell - where one chromosomal region containing the lac operon in E,coli is given, and the other fragment is from a plasmid carrying another lac operon from another source. The two are separated by a slash (/). The possible answers indicate with a ʺ+ʺ or a ʺ-ʺ whether β-galactosidase would be expected to be produced at induced levels under two circumstances: 1) first in the absence of lactose and 2) second in the presence of lactose. (Assume that glucose is not present in the medium.)Genotype F: I+ Oc Z-/ Fʹ I- O+ Z+ KEY:I+ = wild-type repressorI- = mutant repressor (unable to bind to the operator)Is = mutant repressor (insensitive to lactose)O+ = wild-type operatorOc = constitutive operator (insensitive to repressor)arrow_forwardA number of mutations affect the expression of the lac operon in E. coli. The genotypes of several E. coli strains are shown below. ("+" indicates a wild-type gene with normal function and "-" indicates a loss-of-function allele.) Please predict which of the following strains would have the lowest beta-galactosidase enzyme activity, when grown in the lactose medium. Orpt o* z* r* Orpt ot z* Y OrptoztY Orrotzr OrPotz*Yarrow_forwardExplain why (a) inactivation of the O2 or O3 sequence of the lac operon causes only a twofold loss in repression, and (b) inactivation of both O2 and O3 reduces repression ∼70-fold.arrow_forward
- Suppose you have six strains of E. coli. One is wildtype, and each of the other five has a single one of thefollowing mutations: lacZ−, lacY−, lacI−, oc, andlacIS. For each of these six strains, describe thephenotype you would observe using the following assays. [Notes: (1) IPTG is a colorless synthetic molecule that acts as an inducer of lac operon expressionbut cannot serve as a carbon source for bacterialgrowth because it cannot be cleaved byβ-galactosidase; (2) X-gal cannot serve as a carbonsource for growth; (3) E. coli requires active lactosepermease (the product of lacY) to allow lactose,X-gal, or IPTG into the cells.] Colony color in medium containing glycerol as theonly carbon source and X-gal, but no IPTG.d. Colony color in medium containing high levels ofglucose as the only carbon source, X-gal, andIPTG.e. Colony color in medium containing high levels ofglucose as the only carbon source and X-gal, butno IPTGarrow_forwardTrp operon of E. coli is an inducible sytem since it turns on in the presence of tryptophan. In most bacteria, protein synthesis is initiated with a modified methionine residue (N-formylmethionine), whereas unmodified methionines initiate protein synthesis in eukaryotes. Both DNA replication and transcription follow a 5’ to 3’ direction of polarity. Write T if the statement is true and write F if the statement is falsearrow_forwardFor each of the E. coli strains containing lac operon alleles listed, indicate whether the strain is inducible, constitutive, or unable to express beta-galactosidase and permease. (P+ and P- are functional and nonfunctional promoters, respectively) I+ P+ o+ Z- Y+ / I+ P+ oc Z+ Y+ I+ P+ o+ Z+ Y+ / I- P+ oc Z+ Y- I+ P+ o+ Z- Y+ / I- P+ oc Z+ Y- I- P- o+ Z+ Y- / I+ P+ oc Z- Y+ IS P+ o+ Z+ Y+ / I- P+ o+ Z+ Y-arrow_forward
- The lac operon consists of three structural genes, lacZ, lacY and lacA that are transcribed as a single polycistronic mRNA. You are given a new strain of Escherichia coli with the following lac operon genotype: p+0°Z•Y*A +// P*O*Z*Y+ A- (i) Explain how the lac I gene affects gene expression. (ii) Explain the function of the lacP in the bacterial operon. (iii) Which part of the lac operon is cis-dominant? Explain.arrow_forwardTable 1 shows a list of restriction endonucleases with their recognition sequence and the sites of cleavage indicated by arrows. Table 1 Enzyme name Recognition sequence and position of cut 5'GIAATTC3 5'G!GATCC3' 5'GIGTACC3 5'GCIGGCCGC3' 5'IGATC3' 5'GGTACIC3' 5'ALGATCT3 EcoRI ВатHI Аcс651 Notl Sau3A Kpnl BglII (i) Which restriction enzyme(s) produce blunt ends? (ii) Are there any pair of neoschizomers in the list? Explain. (iii) Are there any pair of isocaudomers in the list? Explain.arrow_forwardA number of mutations affect the expression of the lac operon in E. coli. The genotypes of several E. coli strains are shown below. ("+" indicates a wild-type gene with normal function and "-" indicates a loss-of-function allele.) Please predict which of the following strains would have the highest beta-galactosidase enzyme activity, when grown in the lactose medium. O CAP+ r* p* o* z O CAP* I P* o* z* O CAP* r* P O* z* O CAP I P* O z*arrow_forward
- Shown below is a schematic diagram illustrating a very short gene with 5000 bp region of an unknown Schizosaccharomyces pombe genome. (Note: Transcription starts at Transcription Start Site (TSS).) TSS 5. 3' 3 +1 (i) Name the specific regions that can be recognized by Transcription Factor IID (TF ID) and indicate the locations in the diagram above. (ii) List the mechanistic steps that can trigger the initiation of transcription by Transcription Factor IIH (TF IIH).arrow_forwardWhen Zidovudine (3'-azidothymidine or AZT) is added directly to an assay measuring the rate of DNA synthesis catalyzed by the AIDS virus reversed transcriptase, AZT is shown to be without significant inhibitory activity. Explain how AZT is metabolized in virus infected cells to convert the nucleoside into a potent and effective reversed transcriptase inhibitor.arrow_forwardTo characterize the promoter of the gadA operon you made a series of deletion mutants removing pieces of the promoter to see what would happen. The results are found below: gad promoter gada gadX gadz 450 +1 lacz activity transcription start site pH 2.0 pH 7.0 A gad promoter beta-galactosidase (lacZ) +++ 450 gad promoter beta galactosidase (lacZ) +++ +++ 300 +1 gad promoter beta galactosidase (lacZ) 150 D gad beta galactosidase (lacz) -450 150 E gad promoter beta-galactosidase (lacZ) -450 -300 Based on these results, what can you conclude about the gad promoter? O a. The promoter is only regulated by repression Ob. The promoter is regulated by a mix of activation and repression O c. The promoter is only regulated by activation O d. The promoter has multiple operators and multiple enhancersarrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
