
Concept explainers
Can you please help with 1c please
picture with 1 graph is for question 1a)
picture with 4 graphs is for question 1b)
1a) E. coli DNA and binturong DNA are both 50% G-C. If you randomly shear E. coli DNA into 1000 bp fragments and put it through density gradient equilibrium centrifugation, you will find that all the DNA bands at the same place in the gradient, and if you graph the distribution of DNA fragments in the gradient you will get a single peak (see below). If you perform the same experiment with binturong DNA, you will find that a small fraction of the DNA fragments band separately in the gradient (at a different density) and give rise to a small "satellite" peak on a graph of the distribution of DNA fragments in the gradient (see below). Why do these two DNA samples give different results, when they're both 50% G-C?
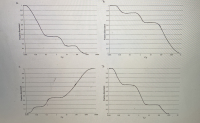
1b) If you denatured the random 1000 bp fragments of binturong DNA that you produced in question 1a by heating them to 95ºC, and then cooled them down to 60ºC and allowed them to reanneal, you would find that approximately 15% of the DNA fragments renatured very rapidly, another 20% of the DNA fragments renatured moderately rapidly, and the remaining DNA fragments renatured relatively slowly. Which of the graphs shown below is a C0t curve showing these results?
1c) What fraction of the binturong genomic DNA would be considered to be "highly repetitive" DNA, based on this data?
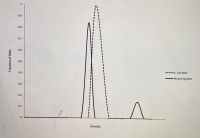
We know that E. coli is a prokaryote and Binturong is a eukaryotic animal. On studying the DNA content of both the organism we see that even though they have the same G-C content, the overall DNA structure is quite different. This is because the prokaryotic genome has a few repetitive DNA sequences whereas repetitive DNA sequences make up a large part of a eukaryotic genome.
Step by stepSolved in 2 steps

- In hybridizing microarrays, we use a solution that contains {3X SSC, 0.5% SDS, 5ug/ml salmon sperm DNA, 5% glycerol}. Describe how to make 100ul of this solution using stocks of 20X SSC, 10% SDS, and 10 mg/ml salmon sperm DNA, sheared, in water. Note that Molecular Biology Grade H2O must be used in making up this solution.arrow_forwardBiology Questionarrow_forwardYou used agarose gel electrophoresis to separate DNA fragments of different size and the experiment worked well. However, you wanted to re run the experiment but this time you made the gel with a higher percentage of Agarose. How might this affect your results compared to the first run? a. There would be no difference between the runs since it is the current, not the agarose that causes migration. b. The higher concentration of agarose would cause the DNA to break apart. c. You can't predict how the concetration of agarose would affect migration. d. The DNA fragments would migrate further down the gel than they did the first time. e. The DNA fragments wouldn't migrate as far down the gel as they did the first time.arrow_forward
- Can you please help with 1c please picture with 1 graph is for question 1a) picture with 4 graphs is for question 1b) 1a) E. coli DNA and binturong DNA are both 50% G-C. If you randomly shear E. coli DNA into 1000 bp fragments and put it through density gradient equilibrium centrifugation, you will find that all the DNA bands at the same place in the gradient, and if you graph the distribution of DNA fragments in the gradient you will get a single peak (see below). If you perform the same experiment with binturong DNA, you will find that a small fraction of the DNA fragments band separately in the gradient (at a different density) and give rise to a small "satellite" peak on a graph of the distribution of DNA fragments in the gradient (see below). Why do these two DNA samples give different results, when they're both 50% G-C? 1b) If you denatured the random 1000 bp fragments of binturong DNA that you produced in question 1a by heating them to 95ºC, and then cooled them down to 60ºC…arrow_forward1) Prepare the following enzymatic reaction, present it in tabulated form. In a final volume of 30 ul, where buffer 4 (10 ml). How much volume of each reagent would be used and how much of water? Is there any problem? 2) The DNA pol 1 enzyme comes at a concentration of 50,000 U/ml. You have to prepare a 50 ug PCR reaction where you must use 0.05 U/ml reaction. You add 10 ul of PCR buffer, 2 ng of tempered DNA that is at a concentration of 0.5 ng/ul, primers (which are at 200 mM) so that each one remains at a concentration of 200 uM, Mg+2 that is 5 mM (10 X), enzyme and water. Present the table of all the reagents included in the reaction, the volumes of each one in ul. Present where the initial and final concentration of each reagent applies. Assume you have micropipettes for all values.arrow_forwardSecond, Question 3arrow_forward
- What is the role of GelRed® in Agarose gel electrophoresis of DNA fragments? Please select the single answer that is most correct GelRed® moves down the agarose gel in response to the electric current and enables visualisation of the position of A the nucleic acids within in the agarose gel. GelRed® intercalates with the Nucleic acid and, under UV light, fluoresces to enable visualisation of the position of the nucleic acids in the agarose gel. GelRed® intercalates with the Nucleic acid and enables visualisation of the position of the nucleic acids in C the agarose gel. GelRed® intercalates with the amino acids in the agarose gel and enables visualisation of the position of their in D the agarose gel.arrow_forwardYou digest 4 uL of plasmid DNA that is 50 ng/uL concetration in a total volume of 20 uL. You run 10 uL of the digest on teh gel. You then do a DNA purification protocol with a Zippy prep on the remaining digested DNA. You elute the DNA in a 25 uL. A 2 uL ssample has the concentration of 2 ng/uL. What is the DNA yield?arrow_forwardCan you please help with 1f please picture with 1 graph is for question 1a) picture with 4 graphs is for question 1b) 1a) E. coli DNA and binturong DNA are both 50% G-C. If you randomly shear E. coli DNA into 1000 bp fragments and put it through density gradient equilibrium centrifugation, you will find that all the DNA bands at the same place in the gradient, and if you graph the distribution of DNA fragments in the gradient you will get a single peak (see below). If you perform the same experiment with binturong DNA, you will find that a small fraction of the DNA fragments band separately in the gradient (at a different density) and give rise to a small "satellite" peak on a graph of the distribution of DNA fragments in the gradient (see below). Why do these two DNA samples give different results, when they're both 50% G-C? 1b) If you denatured the random 1000 bp fragments of binturong DNA that you produced in question 1a by heating them to 95ºC, and then cooled them down to 60ºC and…arrow_forward
- A 2.0kb bacterial plasmid ‘BS1030’ is digested with the restriction endonuclease Sau3A; the plasmid map is depicted in the diagram below and the Sau3A (S) restriction sites are indicated. Which of the following DNA fragments do you expect to see on an agarose gel when you run Sau3A-digested plasmid ‘BS1030’ DNA? a. 250 bp, 450 bp, 550 bp, 1.1 kb, 1.5 kb and 2.0 kb b. 2.0kb c. 250 bp, 400 bp, 450 bp, 500 bp and 550 bp d. 100 bp, 200 bp, 250 bp, 400 bp, 500 bp and 550 bparrow_forwardCan you please help with 1c please picture with 1 graph is for question 1a) picture with 4 graphs is for question 1b) 1a) E. coli DNA and binturong DNA are both 50% G-C. If you randomly shear E. coli DNA into 1000 bp fragments and put it through density gradient equilibrium centrifugation, you will find that all the DNA bands at the same place in the gradient, and if you graph the distribution of DNA fragments in the gradient you will get a single peak (see below). If you perform the same experiment with binturong DNA, you will find that a small fraction of the DNA fragments band separately in the gradient (at a different density) and give rise to a small "satellite" peak on a graph of the distribution of DNA fragments in the gradient (see below). Why do these two DNA samples give different results, when they're both 50% G-C? 1b) If you denatured the random 1000 bp fragments of binturong DNA that you produced in question 1a by heating them to 95ºC, and then cooled them down to 60ºC…arrow_forwardWhich one is the correct one?arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





