
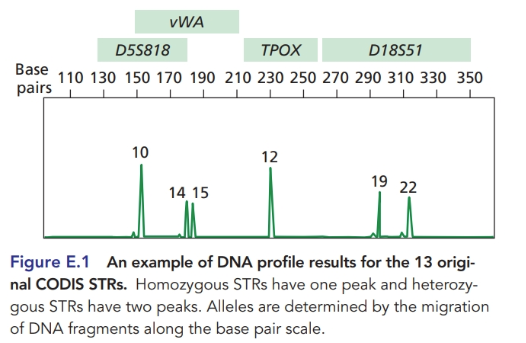
Additional STR allele frequency information can be added to improve the analysis in Problem
E.8 Figure
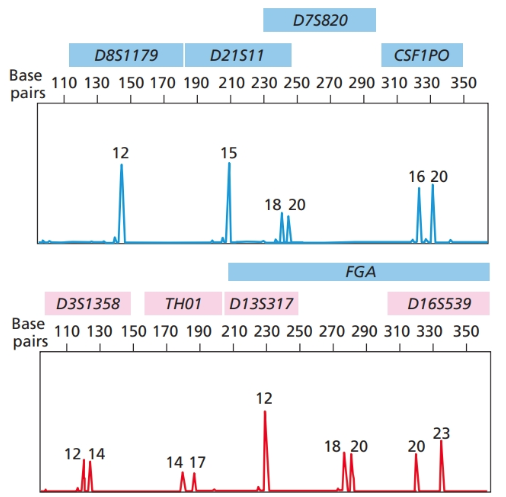

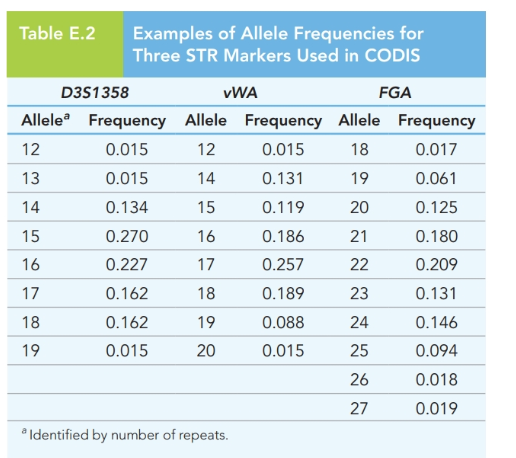
Want to see the full answer?
Check out a sample textbook solution
Chapter E Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Complementation tests of distinct recessive mutants, 1 through 8, produce the data in the matrix below. A plus (+) indicates complementation, meaning the phenotype of the combined alleles is wild type, and a minus (-) indicates a failure to complement meaning that a mutant phenotype results. Assume that the missing mutant combinations would yield data consistent with the entries that are shown. How many complementation groups are formed by these eight mutants? (Picture attached) A) 2 B) 3 C) 4 D) 5 E) 6arrow_forwarda - What is Position-Specific Scoring Matrices (PSSM) and how do we obtain it? b - Construct the HMM for the following sequences where the matches are considered for the columns that don't have any gap T G A C А G T G A А G G T G G A G G C A G EE E E Earrow_forward1. DO NOT ROUND OFF YOUR ANSWERS PREMATURELY. Use the unrounded and complete values when computing for the genetic gain; 2. Upon encoding, all final answers should be rounded-off to TWO decimal places. Examples: 3.5798 = 3.58, 0.0079 = 0.01, 0.00000008 = 0.00; 2. Report your answers as NUMERIC VALUES only. Please exclude the units; 3. Write ONLY one answer for each question. Avoid using "or", "/", "&" and "and". Provide only what is asked to you; 4. AVOID using unnecessary PUNCTUATIONS (. , ? 1; ; etc.), SYMBOLS ( []{ } etc.) and SPACES (before or after your answer). Please follow these instructions carefully. Prediction of Genetic Gain. If the milk yield of a herd of cow is 6,479 kg while those kept for breeding is 6,890 kg, and the h2 estimate of milk yield is 0.85. And the generation interval of dairy cattle is 4 years. What will be the expected genetic gain in the milk yield (kg/year) of the next generation?arrow_forward
- Answer the following questions. 1. Construct a map for the genes d,e,f. Assume that: d and e = 3%; e and f = 5%. Give 2 arrangements of the genes/maps. 2. If d and f = 2%, what is the correct arrangement of the genes d,e,f? 3. Consider the fourth gene "g". if g and e = 1.5%, give two possible arrangements. 4. If d and g = 1.5 % give the correct order of the four genes %3Darrow_forwardB What is the genotype ratio? What is the phenotype ratio? 3. What are the expected genotype and phenotype ratios in the following genetic conditions? Use scratch paper to do the Punnett square if needed but you do not need to draw it on the worksheet. a. Monohybrid cross between 2 heterozygous individuals (Aa x Aa) Genotype ratio: 1:2:1 Phenotype ratio: 3:1 G q 8:32 a b. Dihybrid cross between 2 heterozygous individuals (AaBb x AaBb) Genotype ratio: Phenotype ratio: 4. Both Mrs. Smith and Mrs. Jones had babies the same day in the same hospital. Mrs. Smith took home a baby girl, whom she named Sharon. Mrs. Jones took home a girl, whom she named Jane. Mrs. Jones began to suspect, however, that the child had been accidentally switched with Mrs. Smith baby in the nursery. Blood test were made; Mr. Smith was type A, Mrs. Smith was type B, Mr. Jones was type A, Mrs. Jones was type A. Sharon was type O, and Jane was type. B. Had a mix-up occurred? Use scratch paper, you do not need to…arrow_forwardUse keyboard only to enter your answer below. ALL WORKING MUST BE SHOWN Problem 1) mutation in a gene on chromosome 15 that codes for an enzyme. The disease is an inherited autosomal recessive condition which is Tay-Sachs disease is caused by loss of function found amongst Ashkenazi Jews of Central European origin. In this population, 3 in 5,200 children are born with the disease. What proportion of the population are carriers (heterozygotes) for this disease?arrow_forward
- 3.27 Complementation tests of the recessive mutant g genes a through f produced the data in the accom- panying matrix. The circles represent missing data. beaim Assuming that all of the missing mutant combina- motions would yield data consistent with the entries ithat are known, complete the table by filling each oneel circle with a + or – as needed. - rt yd bat 21 igi a rıb a O + b c d e fi 612 nobsdi b liw insiqans bosale yns mon las 29 ni YOu C ono e 66 BA AA f AAarrow_forwardWhy is the rate of cotransformation for all three genes (M S F) almost the same as the rate of cotransformation for M F alone?arrow_forwardThe DNA of every individual in the pedigree shown in image B (below) has been sequenced at the causative locus, all the non-shaded individuals are wild type apart from III.1 and III.6. III.1 and III.6 have both been proven to have the causative allele for the condition but they do not exhibit any of the phenotypic signs or symptoms. Based on this pedigree, what is the level of penetrance for the condition? Please give your answer as a percentage to one decimal place, give the number only, no percentage symbol. Given the information above I calculate the level of penetrance seen in image B to be "Blank" 1 percent.arrow_forward
- The DNA of every individual in the pedigree shown in image B (below) has been sequenced at the causative locus, all the non- shaded individuals are wild type apart from III.1 and III.6. III.1 and III.6 have both been proven to have the causative allele for the condition but they do not exhibit any of the phenotypic signs or symptoms. Based on this pedigree, what is the level of penetrance for the condition? Please give your answer as a percentage to one decimal place, give the number only, no percentage symbol. ANSWER: Given the information above I calculate the level of penetrance seen in image B to be Blank 1 percent. A KEY Homozygous Homozygous Heterozygous Heterozygous Wild Type Male Female Male Female Male Note: Completely red symbol denotes an individual exhibiting the phenotype of interest CI || III IV V 1/4 1/2 1/2 1/2 1/2 Wild Type Female 1/4 1/2 Affected Known carrier Affected female Normal female Affected male Normal male D ●●●arrow_forwarda - What is Position-Specific Scoring Matrices (PSSM) and how do we obtain it? b - Construct the HMM for the following sequences where the matches are considered for the columns that don't have any gap T G A A G T T G A I A G G C A G T G G C T A Garrow_forwardSearch the menus (Alt+/) 75% Normal text Calibri в IUA 11 + ... 1.| 2 | 3 | • 4.| 5 6 8 12. Although Laurlanthalasa, Princess of the Qualinesti elves, is known for riding into battle on a silver dragon, this is not her main form of transportation. She generally travels by way of griffon. Although the allele for a "crown" of feathers around the head is dominant, it is rarely found in the wild. This may be due to the fact that it makes the griffons stand out, thus making them less effective hunters and more visible to their natural enemies, evil chromatic dragons. This scarcity also makes them prized by elven royalty. Lauralanthalasa was given a breeding pair of bald griffons by an admirer who claims that they are both hybrids for the crowned gene. Create a Punnett Square to determine if Lauralanthalasa will have any chance using these two griffins to produce crowned offspring. Alleles (letters) and Phenotypes All Genotype and Phenotype Parent Punnett Square Possibilities Genotypes…arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





