
Concept explainers
In conducting the study described in Problem
How might you determine whether this set of genes is regulated by a common transcription factor?
How might you approach this question if genome sequences for the related Saccharomyces species, S. paradoxus, S. mikatae, and S. bayanus, were also available?
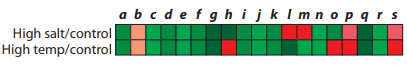
You are studying similarities and differences in how organisms respond to high salt concentrations and high temperatures. You begin your investigation by using microarrays to compare gene expression patterns of S. cerevisiae in normal growth conditions, in
Trending nowThis is a popular solution!

Chapter 16 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Explain how the following mutations would affect transcription of the yeast GAL1 gene in the presence of galactose. (a) A deletion within the GAL4 gene that removes the region encoding amino acids 1 to 100. (b) A deletion of the entire GAL3 gene. (c) A mutation within the GAL80 gene that blocks the ability of Gal80 protein to interact with Gal3p. (d) A deletion of one of the four UASG elements upstream from the GAL1 gene. (e) A point mutation in the GAL1 core promoter that alters the sequence of the TATA box.arrow_forwardSearching the yeast Saccharomyces cerevisiae genome, researchers found approximately 4,000 DNA sites with a sequence which could potentially bind the yeast transcription factor GAL4. GAL4 activates the transcription of galactose genes. Yet there are only 10 GAL4-binding sites which control the genes necessary for galactose metabolism. The GAL4 binding sequence is CGGAT#AGAAGC*GCCG, where # is T, C or G, and * is C or T. In one chromatin immunoprecipitation experiment (ChIP), yeast growing on galactose were lysed, and subjected to cross-linking reagents which cross-linked transcription factors and activators to DNA. Next the DNA was sheared into small fragments, and antibodies to GAL4 were added. These antibodies coprecipitated the GAL4 and the DNA it was cross-linked to. The cross-linking was then chemically reversed, and the DNA was isolated, cloned into a library of plasmids and sequenced. Results showed that only 10 different DNA sequences had GAL4 bound. Since the…arrow_forwardAssume that a series of compounds has been discovered in Neurospora. Compounds A–F appear to be intermediates in a biochemical pathway. Conversion of one intermediate to the next is controlled by enzymes that are encoded by genes. Several mutations in these genes have been identified and Neurospora strains 1–4 each contain a single mutation. Strains 1–4 are grown on minimal media supplemented with one of the compounds A–F. The ability of each strain to grow when supplemented with different compounds is shown in the table (+ = growth; o = no growth). Which biochemical pathway fits the data presented? Media Supplement Strain A B C D E F 1 o o o + + + 2 o o o o + + 3 o o o o + o 4 o o + + + + A) A → B → C → D → E → F B) A → B → C → F → D → E C) F → B → C → D → A → E D) A → B → C → D → F → E E) A → B → F → E → C → Darrow_forward
- Describe how transcription would be affected in the Galactose metabolizing pathway in Yeast in the presence of the following mutations. 1. A mutation that resulted in an inability of Gal80 to enter the nucleus. 2. A mutation that resulted in a lack of ability of Gal3 to bind galactose.arrow_forwardThe entire genome of the yeast Saccharomyces cerevisiaehas been sequenced. This sequencing has led to the identification of all the open reading frames (ORFs, gene-sizesequences with appropriate translational initiation andtermination signals) in the genome. Some of these ORFsare previously known genes with established functions;however, the remainder are unassigned reading frames(URFs). To deduce the possible functions of the URFs,they are being systematically, one at a time, convertedinto null alleles by in vitro knockout techniques. The results are as follows:15 percent are lethal when knocked out.25 percent show some mutant phenotype (alteredmorphology, altered nutrition, and so forth).60 percent show no detectable mutant phenotype at alland resemble wild type.Explain the possible molecular-genetic basis of thesethree mutant categories, inventing examples wherepossible.arrow_forwardThe locations of the TATA box in two species of yeast, Saccharomyces pombe and Saccharomyces cerevisiae, differ dramatically. The TATA box of S. pombe is about 30 nucleotides upstream of the transcription start site, similar to the location in most other eukaryotic cells. However, the TATA box of S. cerevisiae is 40 to 120 nucleotides upstream of the start site. To better understand what sets the start site in these organisms, researchers at Stanford University conducted a series of experiments to determine which components of the transcription apparatus of these two species could be interchanged (Y. Li et al. 1994. Science 263:805–807). In these experiments, different general transcription factors and RNA polymerases were switched in S. pombe and S. cerevisiae, and the effects of each switch on the level of RNA synthesis and on the starting point of transcription were observed. The results from one set of experiments are shown in the table below. Components cTFIIB, cTFIIE, cTFIIF,…arrow_forward
- The locations of the TATA box in two species of yeast, Saccharomyces pombe and Saccharomyces cerevisiae, differ dramatically. The TATA box of S. pombe is about 30 nucleotides upstream of the transcription start site, similar to the location in most other eukaryotic cells. However, the TATA box of S. cerevisiae is 40 to 120 nucleotides upstream of the start site. To better understand what sets the start site in these organisms, researchers at Stanford University conducted a series of experiments to determine which components of the transcription apparatus of these two species could be interchanged (Y. Li et al. 1994. Science 263:805–807). In these experiments, different general transcription factors and RNA polymerases were switched in S. pombe and S. cerevisiae, and the effects of each switch on the level of RNA synthesis and on the starting point of transcription were observed. The results from one set of experiments are shown in the table below. Components cTFIIB, cTFIIE, cTFIIF,…arrow_forwardThe locations of the TATA box in two species of yeast, Saccharomyces pombe and Saccharomyces cerevisiae, differ dramatically. The TATA box of S. pombe is about 30 nucleotides upstream of the transcription start site, similar to the location in most other eukaryotic cells. However, the TATA box of S. cerevisiae is 40 to 120 nucleotides upstream of the start site. To better understand what sets the start site in these organisms, researchers at Stanford University conducted a series of experiments to determine which components of the transcription apparatus of these two species could be interchanged (Y. Li et al. 1994. Science 263:805–807). In these experiments, different general transcription factors and RNA polymerases were switched in S. pombe and S. cerevisiae, and the effects of each switch on the level of RNA synthesis and on the starting point of transcription were observed. The results from one set of experiments are shown in the table below. Components cTFIIB, cTFIIE, cTFIIF,…arrow_forwardWhat percentage of the DNA sites in yeast are accessible, assuming that the fraction of sites observed for GAL4 is typical? To how many base pairs of the 12-Mb yeast genome does this percentage correspond?arrow_forward
- Let’s suppose you make a transposon library of the cellulose-secreting bacterium Komagataeibacter xylinus, with the goal of finding mutants that produce higher than normal amounts of cellulose, which would be useful industrially. However, despite your best efforts you are unable to isolate any transposon mutants that make more cellulose than the wild-type strain.Why might this have failed? List as many reasons as you can think of.arrow_forwardPolymerase Chain Reaction (PCR) was invented by Kary Mullis in 1983. This technique had indeed facilitated research in various areas of molecular biology and genetics. You would like to amplify a particular gene fragment from the yeast genome using Polymerase Chain Reaction (PCR): (i) What is the major enzyme required to kick start this reaction? (ii) Besides the enzymes mentioned in Q2 (a)(), state three (3) other chemicals required for the amplification process to be carried out.arrow_forwardFor the lac genotypes of Escherichia coli shown in the following Table 1, predict the expression of beta-galactosidase (Z) and permease (Y) is inducible or noninducible or constitutive. Explain your answer. Table 1 Genotype I-P+O+Z+Y+ (i) (ii) I+P+OCZ+Y+ (iii) ISP+O+Z+Y+ (iv) I+P+O+Z+Y-//I+P-O+Z+Y+ (v) ISP+OcZ+Y+//I-P+O+Z+Y- Condition No lactose No lactose lactose lactose No lactosearrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





