
Concept explainers
A muscle enzyme called ME
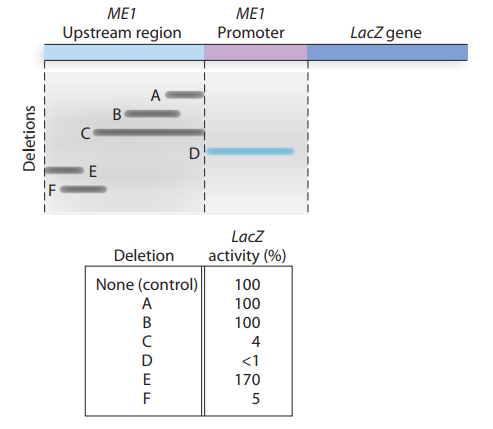
a. Does this information indicate the presence of enhancer and
b. Why does deletion D effectively eliminate transcription oflacZ?
c. Given the information available from deletion analysis, can you give a molecular explanation for the observation that ME
Want to see the full answer?
Check out a sample textbook solution
Chapter 13 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- The following diagram show what is required for an active promoter of a gene of interest, where:A1 = Activator 1A2 = Activator 2Med = MediatorRep = Repressor Based on the following data, predict: Chromatin conformation Methylation state of the proximal promoter If protein A1 is present or absent If protein A2 is present or absent If the mediator is present or absent If the repressor is absence or present If this gene is likely to be transcribed or notPlease note that this is an "all or nothing" bonus question, and no partial credit will be awarded. Selecting all answers will result in zero points awarded. Selecting at least one incorrect answer will result in zero points being awarded. Question 28 options: Euchromatin Heterochromatin Methylated promoter Unmethylated promoter Activator A1 present Activator A1 absent Activator A2 present…arrow_forwardDiscuss the following argument: “if the expression of every gene depends on a set of transcription regulators, then the expression of these regulators must also depend on the expression of other regulators, and their expression must depend on the expression of still other regulators, and so on. cells would therefore need an infinite number of genes, most of which would code for transcription regulators.” how does the cell get by without having to achieve the impossible?arrow_forwardThe yeast gene SER3, whose product has a role in serine biosynthesis, is repressed during growth in nutrient-rich medium, so little transcription takes place, and little SER3 enzyme is produced, under these conditions. In an investigation of the repression of the SER3 gene, a region of DNA upstream of SER3 was found to be heavily transcribed when SER3 is repressed ). Within this upstream region is a promoter that stimulates the transcription of an RNA molecule called SRG1 RNA (for SER3 regulatory gene 1). This RNA molecule has none of the sequences necessary for translation. Mutations in the promoter for SRG1 result in the disappearance of SRG1 RNA, and these mutations remove the repression of SER3. When RNA polymerase binds to the SRG1 promoter, the polymerase travels downstream, transcribing the SGR1 RNA, and passes through and transcribes the promoter for SER3. This activity leads to the repression of SER3. Propose a possible explanation for how the transcription of SGR1 might…arrow_forward
- Regulation of transcription is mediated by proteins that first must bind to specific sequences/elements that are present in DNA. Fill in the table below based on your knowledge of the different sequences and proteins involved in the regulation of eukaryotic gene expression. Regulatory sequence Category of Protein that Binds Effect on Transcription of Associated Gene Generic or Gene-Specific Regulation? Core Promoter Element Activator Silencerarrow_forwardGal4 is a transcription factor that activates transcription of galactose metabolism genes in yeast. These genes are ‘turned on’ when yeast cells need to metabolize galactose. To identify promoter sequences necessary for regulation of transcription of GAL1, reporter gene fusions were made and introduced into yeast cells. Deletions of GAL1 promoter were cloned upstream of LacZ gene. β-Galactosidase activity was measured in presence of galactose. Shown below is a representation of the results obtained. In the diagrams below (not to scale!): • Construct 1 contains ~ 130bp of the promoter, which is predicted to have all the predicted/putative proximal promoter elements (indicated by the solid boxes) needed to regulate transcription of GAL1.• The stippled box is the core promoter.• The arrow represents the transcriptional start site for the reporter gene Lac Z• Number of + signs represents level of transcription• Star represents a mutation in DNA sequence at that location (few nucleotides…arrow_forwardIn the sea urchin, early development may occur even in the presence of actinomycin D, which inhibits RNA synthesis. However, if actinomycin D is present early in development but is removed a few hours later, all development stops. In fact, if actinomycin D is present only between the sixth and eleventh hours of development, events that normally occur at the fifteenth hour are arrested. What conclusions can be drawn concerning the role of gene transcription between hours 6 and 15?arrow_forward
- Negative supercoiling of DNA favors the transcription of genes because it facilitates unwinding. However, not all promoter sites are stimulated by negative supercoiling. The promoter site for topoisomerase II itself is a noteworthy exception. Negative supercoiling decreases the rate of transcription of this gene. Propose a possible mechanism for this effect and suggest a reason why it may occur.arrow_forwardConsider this list (below) of steps involved in transcription. These steps are out of order. TRANSCRIPTION: 1. mRNA travels through a nuclear pore and enters the cytoplasm 2. the mRNA polymerase attaches at the start of a specific gene 3. RNA polymerase reads the gene surface4. a transcription factor bonds to a promoter site5. DNA molecule is unwound 6. a complimentary mRNA is produced What is the correct order of this transcription?arrow_forwardCD3 is a signaling protein that is typically found only in the plasma membrane of immune system T lymphocytes. CD3 is composed of several different polypeptides, including a gamma chain, CD3γ. Scientists analyzed the promoter of the CD3γ chain gene for regulatory sequences that might have positive or negative effects on expression of the gene. The scientists cloned fragments of the CD3γ gene that included the first transcribed nucleotides plus up to 789 nucleotides of upstream regulatory sequences into plasmids in which the gene for the firefly enzyme luciferase immediately follows the fragments. The plasmids were then introduced into a line of T lymphocytes (Figure 1), and the cells were allowed to grow for a short while. Because the regulatory sequences of the CD3γ gene immediately precede the luciferase gene in the plasmids, the activity, either positive or negative, of the regulatory sequences affected the amount of luciferase gene expression by the T lymphocytes. Luciferase…arrow_forward
- Progesterone is a steroid hormone (also described as a ligand) that prepares the body for pregnancy. It binds to the progesterone receptor (PR) protein in the cytoplasm of various cells. Ligand bound PR acts as a transcriptional activator, binds to the DNA in the promoter region of several genes and leads to transcriptional activation of these genes. Ligand bound PR has been shown to increase the expression of a gene, FKBP5. You are studying the activity of wild-type (WT) and mutant PR in cells by examining expression of FKBP5. Results are obtained as shown in the figure below, where the asterisk indicates when progesterone was (or was not) added to the cells. From the results, which of the following statements can be concluded? WT PR without progesterone WT PR with progesterone Time Time mutant PR without progesterone mutant PR with progesterone Time Time The wild-type PR is unable to increase FKBP5 expression in the absence of ligand The wild-type PR increases FKBP5 expression after…arrow_forwardA strain of Arabidopsis thaliana possesses a mutation in the APETALA2 gene. As a result of this mutation, much of the 3′ UTR of the mRNA transcribed from the gene is deleted. What is the most likely effect of this mutation on the expression of the APETALA2 gene?arrow_forwardIIVY is a LysR regulator, and it helps control transcription of a neighboring gene, ilvC. IlvY can either bend the DNA and block RNAP's access to ilvC's -35 promoter site, or it can allow access to the ilvC -35 site, thereby promoting ilvC transcription. What determines whether llvY is repressing or activating transcription from the ilvC promoter? The presence of the co-inducer acetolactate - acetolactate binds to llvY and changes llvY's shape such that it doesn't block the -35 of the ilvC promoter. IlvY represses ilvC transcription when the IlvC protein is bound to llvY, and it activates ilvC transcription when the lvC protein is not present to bind to llvY. The presence of the co-inducer valine - when valine is present, it binds llvY and causes llvY to repress transcription from the ilvC promoter. The presence of the co-inducer acetolactate - acetolactate induces negative supercoiling at the ilvC promoter, which represses ilvC transcription.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





