
Concept explainers
You have identified a
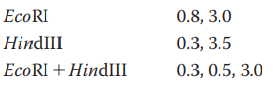
Trending nowThis is a popular solution!

Chapter 15 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Below is a diagram of the vector you are planning to use. You identify four restriction enzyme recognition sites in the vector as indicated in the diagram below. The distance between the restriction sites are indicated by numbers (in kilo base pairs). ori ВатHI ВатHI 2 ЕcoRI Kpnl If you digest the vector with a combination of EcoRI and BamHI and run the resulting DNA fragments on a gel, which lane would represent the expected result? (Lane 1 contains the DNA size marker.) А В с (kbp) 8 Promoterarrow_forwardYou have isolated a cDNA clone encoding a protein of interest in a higher eukaryote. This cDNA clone is not cleaved by restriction endonuclease EcoRI. When this cDNA is used as a radioactive probe for blot hybridization analysis of EcoRI-digested genomic DNA, three radioactive bands are seen on the resulting Southern blot. Does this result indicate that the genome of the eukaryote in question contains three copies of the gene encoding the protein of interest? Explain.arrow_forwardThe figure below shows the recognition sequences and cleavage positions of three restriction enzymes.You plan to ligate DNA from two different sources. The target DNA is digested with BamHI,and the insert DNA is digested with BglII, and the resulting fragments mixed and incubatedwith DNA ligase. a) Write out the sequence (in double-stranded format) of the longest insert fragment that will result after BglII digestion, ensure the nature of the overhangs is clear.b) Write out the sequence (in double-stranded format) of the ligation product, with the insert fragment joined into the BamHI site of the target DNA. Use black for target sequences, and blue for insert sequences. c) Assume the ligation reaction was successful and you have generated a recombinant DNAmolecule. Which of the three enzymes listed above can be used to excise the insert DNAfrom the target? Motivate your answer.arrow_forward
- After characterizing the DNA composition of various cats, you identify a protein-coding gene in tigers called stripes and wish to study the structure of the protein product STRIPES. This requires that you purify recombinant ridges from E. coli. First, the stripes gene must be amplified by PCR and then inserted into an appropriate plasmid for bacterial expression. Such a plasmid is diagrammed below. ori CAP Binding Site من Promoter MCS The restriction sites for Aatll and Kpnl are: Aatll 5'-GACGTC-3' Kpnl = 5'-GGTACC-3' Laco (Operator) -Kpnl Aatll The coding strand for the stripes gene is shown below, with start and stop codons in bold. 5'-ATGCAACAGTAGCTGAAGCCCAGTGACACCATCGAAAATGTGAAGGCCAAGATGAGGCTCATCTTTGCAGGCAAGCAGCTG GAAGATGGCCGTACTCTTTCTGACTATGCGTCTGAGAGGTGGTATGCAGATCTTCGTGAAGACCCTGACCGGCAAGACCAATGT GAAGGCCAAGATCCAGGATAAAGAAGGCATCCCTCCCGACCAGCAGAGGGCACTCTTTCTGACTACAACATCCAGAAGGAGTCG ACCCTGCACCTGGTCCTGCTGACCGGCAAGACCATCACTCTGGAGGTGGAGCCCAGTGACACCATCGAAAATCCCGACCAGCAG…arrow_forwardKnowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardWhen the restriction endonuclease EcoRI is used to digest a 10 kb DNA fragment, it produces 4 kb and 6 kb-sized fragments. Digesting the 10 kb fragment with BamHI yields three fragments, each ranging in size from one to three and a half kilobytes. Four pieces of 0.5, 1, 3 and 5.5 kb are formed after using both enzymes. Create a restriction map for this 10 kb piece of DNA using the information you have collected. Make a note of where the two enzymes cut, as well as the distances between the enzymes.arrow_forward
- A cloned Bam HI DNA fragment, 3.0 kb long, was isolated in preparation for sequence determination. First, a map of restriction endonuclease cleavage sites had to be prepared. Cleavage by each of the indicated enzymes generated fragments of the sizes shown on the three agarose electrophoresis gel figures below; the gels are running from top to bottom. On the diagram below those, show the relative positions of EcoR I and Hpa II cleavage sites and the distances (in kb) between them. Explain your reasoning – is your placement of the cleavage sites the only one that will work?arrow_forwardYou have a 5.3 kb PstI fragment cloned into the PstI site of the vector pUC19, which is 2.7 kb in size. This vector has unique sites for the following enzymes in a multiple cloning site: PstI, HincII, XbaI, BamHI, SmaI, EcoRI A restriction map of the 5.3 kb insert is prepared. The recombinant plasmid is digested with the enzymes listed above in single digests, and then several combinations of enzymes are tested in double digests. The following bands are observed when the digests are run on a gel: Use the data above to construct a map of the cloned insert. Note that fragments smaller than 100 bp will not usually be visible on a gel, and that 2 fragments with a similar size will run at the same spot on a gel – resulting in a brighter band.arrow_forwardTransposon mutagenesis was used to generate a library of mutants within the Salmonella genome. You are trying to identify a colony with the transposon inserted in the pathogenic related gene SPI-1 using PCR. Forward and reverse primers are generated that flank either side of the gene and yield a wild type product that is 900 bases in length. Which of the colonies sampled in the gel would you expect to contain the SPI-1 gene with transposon insertion? 3,000 2,000 1,000 700 500 300 100 Ladder Colony A Colony B Colony C Colony D Colony E none colonies A&C colonies B&E O colonies A, C, &D colonies B, D, &E -arrow_forward
- Restriction endonucleases are bacterial enzymes that cleave duplex (double-stranded) DNA at specific nucleotide sequences. The mode of replication of the animal virus SV40 has been investigated by using restriction endonucleases that cleave SV40 DNA into a number of unique segments. Like most viruses, SV40 DNA is circular. The map positions of the 11 fragments produced by a pair of restriction endonucleases are shown on the next page. Immediately following a 5 or 10 minute pulse of radioactively labeled thymidine, labeled SV40 molecules that have completed replication during the pulse are isolated. These newly replicated DNA molecules are digested by the restriction endonucleases and the resulting fragments are analyzed for the relative amounts of pulse label they contain. The results are in the table below. Assume that at the time the label was added there was a random population of replicating SV40 DNA molecules in all possible stages of synthesis. From the information given below,…arrow_forwardThe plasmid cloning vector pBR322 is cleaved with the restriction endonuclease PstI. An isolated DNA fragment from a eukaryotic genome (also produced by PstI cleavage) is added to the prepared vector and ligated. The mixture of ligated DNAs is then used to transform bacteria, and plasmid-containing bacteria are selected by growth in the presence of tetracycline. The cloned DNA fragment is 1,000 bp long and has an EcoRI site 250 bp from one end. Three different recombinant plasmids are cleaved with EcoRI and analyzed by gel electrophoresis, giving the patterns shown below. What does each pattern say about the cloned DNA? Note: pBR322, the PstI and EcoRI restriction sites are about 750 bp apart. The entire plasmid with no cloned insert is 4,361 bp. Size markers in lane 4 have the number of nucleotides noted.arrow_forwardb. After molecular cloning and confirming you properly cloned stripes into the vector shown, you transform bacteria with the plasmid and select out bacteria that contain the plasmid. Now, you want to these transformed bacteria to express the STRIPES. Under what conditions would we need to grow bacteria transformed with this plasmid so that they express STRIPES? Explain your answer, and be sure to describe the important regulatory regions on the plasmid above and what interacts with these regulatory regions in the conditions you have indicated.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





