
Concept explainers
Three recessive traits in garden pea plants are as follows: yellow pods are recessive to green pods, bluish green seedlings are recessive to green seedlings, creeper (a plant that cannot stand up) is recessive to normal. A true-breeding normal plant with green pods and green seedlings was crossed to a creeper with yellow pods and bluish green seedlings. The
2059 green pods, green seedlings, normal
151 green pods, green seedlings, creeper
281 green pods, bluish green seedlings, normal
15 green pods, bluish green seedlings, creeper
2041 yellow pods, bluish green seedlings, creeper
157 yellow pods, bluish green seedlings, normal
282 yellow pods, green seedlings, creeper
11 yellow pods, green seedlings, normal
Construct a genetic map that indicates the map distances between these three genes.
To review:
A genetic map indicating map distances between the genes for pod color, seedling color, and plant stature.
Introduction:
Thepair of alleles ofa gene determines the protein encoded by the genes, and results in the phenotype of the trait. True breeders have the monomorphic condition for the alleles of a trait. A genetic map helps in describing the arrangement of the genes on a particular chromosome.
Explanation of Solution
A normal pea plant, which is truebreeding for green seedling and green pods is crossed with a creeper pea plant, having bluish-green seedlings and yellow pods. The F1 offspring are then crossed with a creeper having bluish-green seedlings and yellow pods.
Different characters of the pea plant can be denoted as follows:
Dominant characters: G for green pods, S for green seedling, and C for normal plants
Recessive characters: g for yellow pods, s for bluish green seedlings, and c for the creeper.
Given, the true breeding normal plant, having green seedlings as well as green pods is crossed with true breeding creeper, having bluish green seedling and yellow pods. The genotype of true breeding plants is GGSSCC and ggsscc. Thus, the gametes will be GSC and gsc. The plants in the F1 generation obtained by crossing these true breeders will be GgSsCc.
When three genes are linked then G, S, and C alleles will be linked together whereas g, s, and c alleles will be linked together, on a homologous chromosome. Given, the F1 plants GgSsCc are crossed with ggsscc (creepers having yellow pods and bluish green seedlings) and following results were obtained:
| Number of plants | Phenotype |
| 2059 | Green pods, green seedlings, normal |
| 151 | Green pods, green seedlings, creepers |
| 281 | Green pods, bluish green seedlings, normal |
| 15 | Green pods, bluish green seedlings, creepers |
| 2041 | Yellow pods, bluish green seedlings, creepers |
| 157 | Yellow pods, bluish green seedlings, normal |
| 282 | Yellow pods, green seedlings, creepers |
| 11 | Yellow pods, green seedlings, normal |
The genetic map for the three genes can be constructed by using the below-mentioned formula for map distance:
The distance between the three genes can be measured by separating the data of the offspring and phenotypic categories into gene pair, and then calculating the map distance between the two genes.
For the map distance between the genes for plant stature and pod color,
the number of offsprings is calculated for each pair of plant stature and color of pods.
Non-recombinant offspring are 2340 normal, green pods, and 2323 creeper, yellow pods.
Recombinant offspring are 166 creeper, green pods, and 168 normal, yellow pods.
Therefore, the map distance will be calculated as,
For the map distance between the genes for plant stature and seedling color, the number of offspring having genes for plant stature and color of the seedlingis given as follows:
Nonrecombinant offspring are 2070 normal, green seedlings, and 2056 creeper, bluish green seedlings, and the recombinant offspring are 433 creeper, green seedlings, and 438 normal, bluish green seedlings.
Therefore, the map distance will be calculated as.
For the map distance between the genes for seedling color and pod color, the number of offspring having genes for the color of seedling and pods is given as follows:
Non-recombinant offspring are 2210 green seedling, green pods, and 2198 bluish green seedling, yellow pods, and the recombinant offspring are 296 bluish green seedling, green pods, and 293 green seedling, yellow pods.Therefore, the map distance will be calculated as,
Thus, the distance between the genes for seedling and pods colors is 11.8 mu, and that between the genes for plant stature and seedling color is 17.4 mu and for the genes for plant stature and pod color, it is 6.7 mu. Therefore, the order of the three genes, according to the map distance in-between them will be seedling color, pod color, plant stature. Gene for pod color is present in the middle. Genetic map is shown below:
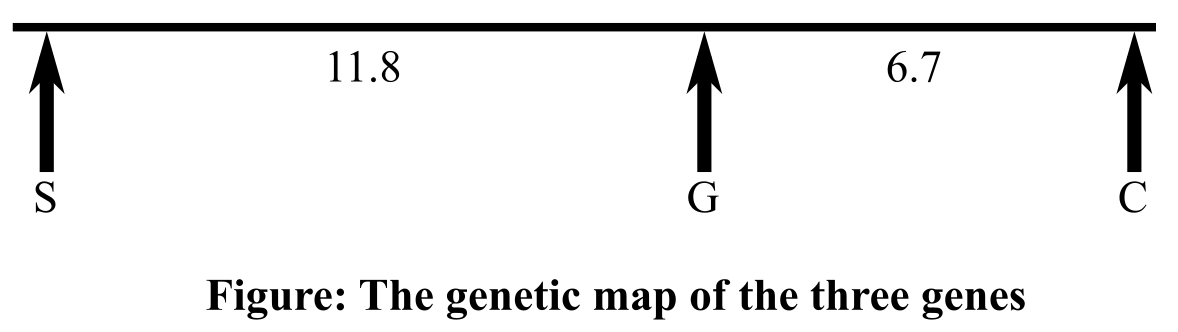
Therefore, it can be concluded that according to the map distance calculated in-between the genes for pod color, plant stature, and seedling color, the order of the three genes will be seedling color, pod color, plant stature(or the opposite order). The distance between the genes responsible for seedling and pods colors is 11.8 mu, and between the genes for plant stature and pod color, it is 6.7 mu. The genes that code for plant stature and seedling color are 17.4 mu apart.
Want to see more full solutions like this?
Chapter 6 Solutions
GENETICS:ANALYSIS+PRIN.(LL)-W/ACCESS
- In pea plants, seed shape and seed color are controlled by genes located on different chromosomes. Seeds may be round (R) or wrinkled (r), with the allele for round seeds being dominant. Alleles for seed color are yellow and green, with the green allele (y) recessive to the yellow (Y) allele. If you cross an individual that is homozygous round and yellow with an individual that is homozygous for wrinkled and green, what is the genotype of the F1 individuals? Set up a Punnett square for the dihybrid cross.arrow_forwardIn corn, two independent, recessive nuclear genes, japonica (j) and iojap (ij), produce variegation (green and white striped leaves). Matings between individuals heterozygous for japonica always produce 3 green:1 striped individuals regardless of how the cross is performed. You have a variegated plant that could be either jj or ijij . What cross can you make to determine the genotype of this plant, and what results do you expect in the F1 generation in each case?arrow_forwardGregor Mendel examined the inheritance of two traits in pea plants: seed coat texture and colour. Seed coat texture can be represented as S-smooth and s-wrinkled, and seed coat colour can be represented as Y-yellow and y-green. SSYY plants were crossed with ssyy plants to yield F1 pea seeds that were all smooth and all yellow. By crossing plants grown from these F1 seeds, Mendel obtained four different phenotypes of F2 seeds: • smooth and green seeds wrinkled and green seeds smooth and yellow seeds wrinkled and yellow seeds ● Use the following information to answer the next question. ● The F2 phenotypic ratio that Mendel obtained upon crossing two heterozygous smooth and yellow F1 individuals would have been: smooth and green wrinkled and green : smooth and yellow: wrinkled and yellow Record only the numeric values associated with the phenotypes. (Do not include the colons, spaces, commas, etc.)arrow_forward
- In corn plants, a dominant allele (K) allows kernel colour and a recessive allele (k) inhibits kernel colour when homozygous. On a different chromosome, the dominant gene P causes purple kernel colour and the homozygous recessive genotype causes red kernel colour.A true breeding white corn plant was crossed with a purple corn plant, yielding 50% red corn plants and 50% purple corn plants.What are the genotypes of the parental corn plants? Select one: a. KKPp kkpp b. KkPP kkPP c. kkPp KkPp d. KKPP kkPparrow_forwardIn the Pea plant, tall plant height (T) is dominant over short (t). Pure-breeding tall and short plants are crossed. A) If the F1 is self-crossed and 400 F2 plants are raised, how many would be expected in each phenotypic class? B) How many of the F2 would be expected to be pure breeding when selfed?arrow_forwardA certain species of morning glories produces flowers that are blue, red, or purple. Two pure-breeding purple lines are crossed and produce F1 progeny that all make blue flowers. The F1 are allowed to self and produce 320 F2 progeny with the following distribution: 185 blue, 115 purple, and 20 red. Which the following is NOT consistent with this information? A) Red-flowering plants are homozygous recessive for both genes. B) The pure-breeding parental parents are homozygous recessive for mutations in two different genes. C) Dominant gene interaction appears to result in a 9:6:1 ratio. D) Blue-flowering plants are either A_bb or aaB_. E) Analysis of the F1 and F2 progeny phenotypes suggests epistasis.arrow_forward
- In the tomato plant, red fruit (R) is dominant over green (r). Smooth fruit skin (S) is dominant over peachy skin (s). The genes for fruit colour and fruit texture are linked on the same chromosome. A heterozygous red, heterozygous smooth plant was crossed with a green, peachy plant. The results are given:Smooth-red = 28Smooth-green 567Peachy-red = 534Peachy-green = 34i. Calculate the map distance between the genes.ii. The map distance calculated in (i) is not representing the true distance between the genes? Provide your argument.arrow_forwardYou have a purple-flowered pea plant, but you do not know if it is homozygous (PP) or heterozygous (Pp) for flower color because both genotypes result in the same purple phenotype. Purple color allel (P) is dominant over white flower allel (p). What would you do to determine the genotype of flower color of this plant? Lötfen birini seçin: O a. Crossing the plant with homozygous large flowered pea plant (LL) Ob. Crossing the plant with heterozygous purple flowered pea plant (Pp) Crossing the plant with homozygous dominant purple flowered pea plant (PP) d. Crossing the plant with a plant whose genotype is unknown e. Crossing the plant with homozygous recessive white flowered pea plant (pp)arrow_forwardA pea plant has a mendilian completely dominant gene for height flower colour phenotype. A pure breeding tall purple flower plant is crossed with a white dwarf plant. All offspring are purple and tall. If one of these offspring are crossed again with the white dwarf plant, what is the chances of producing a tall white flower? Explain your answer and any assumptions that you had to make. Note: You may not need the entire punnett square below or you may wish to use a different method.arrow_forward
- In watermelons, bitter fruit (B) is dominant over sweet fruit (b), and yellow spots (S) are dominant over no spots (s). The genes for these two characteristics assort independently. A homozygous plant that has bitter fruit and yellow spots is crossed with a homozygous plant that has sweet fruit and no spots. The F1 are intercrossed to produce the F2. What will be the phenotypic ratios in the F2? 2. If an F1 plant is backcrossed with the bitter, yellow-spotted parent, what phenotypes and proportions are expected in the offspring? 3. If an F1 plant is backcrossed with the sweet, non-spotted parent, what phenotypes and proportions are expected in the offspring? 4. In cats, curled ears (Cu) result from an allele that is dominant over an allele for normal ears (cu). Black color results from an independently assorting allele (G) that is dominant over an allele for gray (g). A gray cat homozygous for curled ears is mated with a homozygous black cat with normal ears. All the F1…arrow_forwardIn Bean plants two genes are defined below with the lower case letter indicating the recessive allele. yellow flower color a = green flower color. B = spotted pods b = plain pods A = A parental generation cross is made between a pure breeding plant with yellow flower color and spotted pods, to a pure breeding plant with green flower color and plain pods. Write this parental generation cross using the symbols defined above. Write the genotype of the F1 plants. An F1 plant is then testcrossed. Write this testcross using the symbols defined above.arrow_forwardConsider this cross in pea plants: Tt Rr yy Aa × Tt rr Yy Aa, whereT = tall, t = dwarf, R = round, r = wrinkled, Y = yellow, y = green,A = axial, a = terminal. What is the expected phenotypic outcomeof this cross? Have one group of students solve this problem bymaking one big Punnett square, and have another group solve it bymaking four single-gene Punnett squares and using the multiplication method. Time each other to see who gets done first.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





