
Concept explainers
How does DNA replication occur in a precise manner to ensure that identical genetic information is put into the new chromatid? See Figures 8.12 and 8.13.
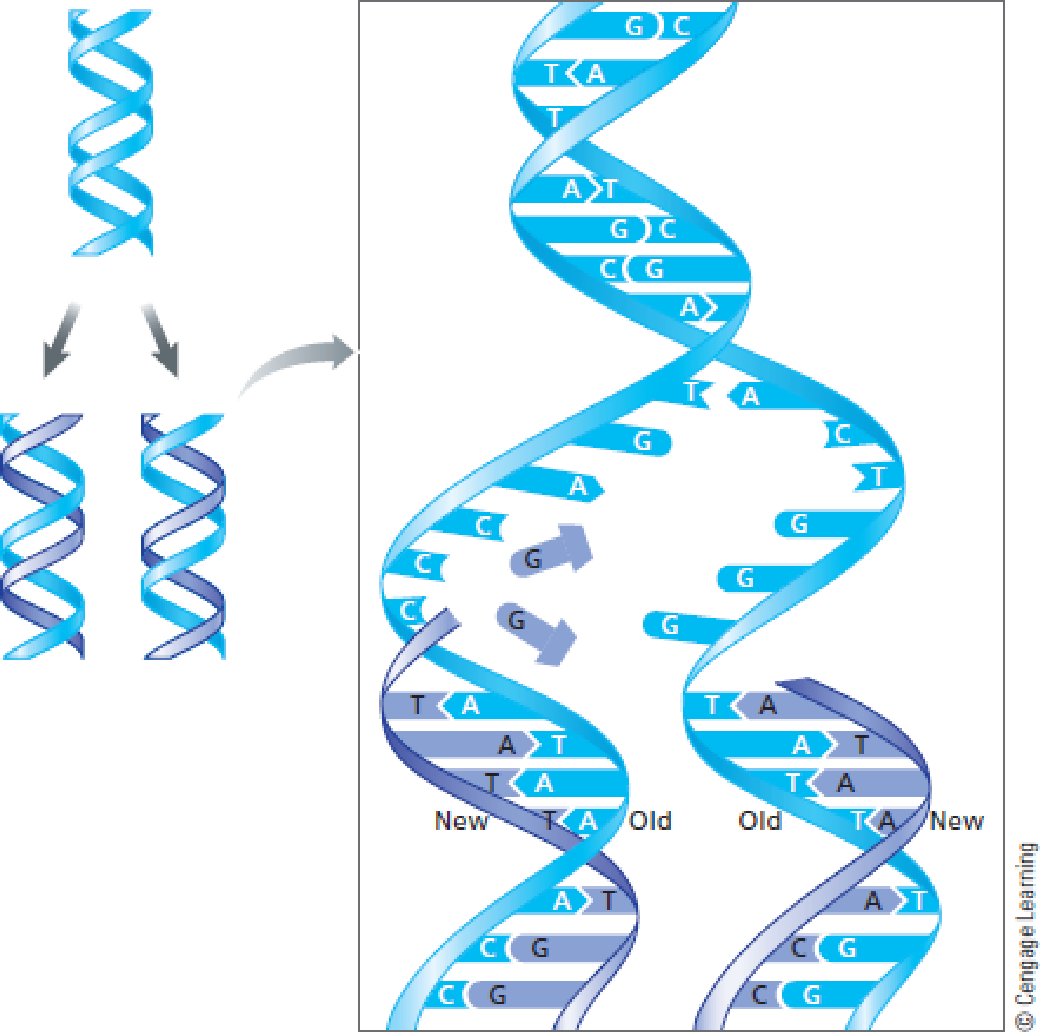
FIGURE 8.12 In DNA replication, the two polynucleotide strands uncoil, and each is a template for synthesizing a new strand. A
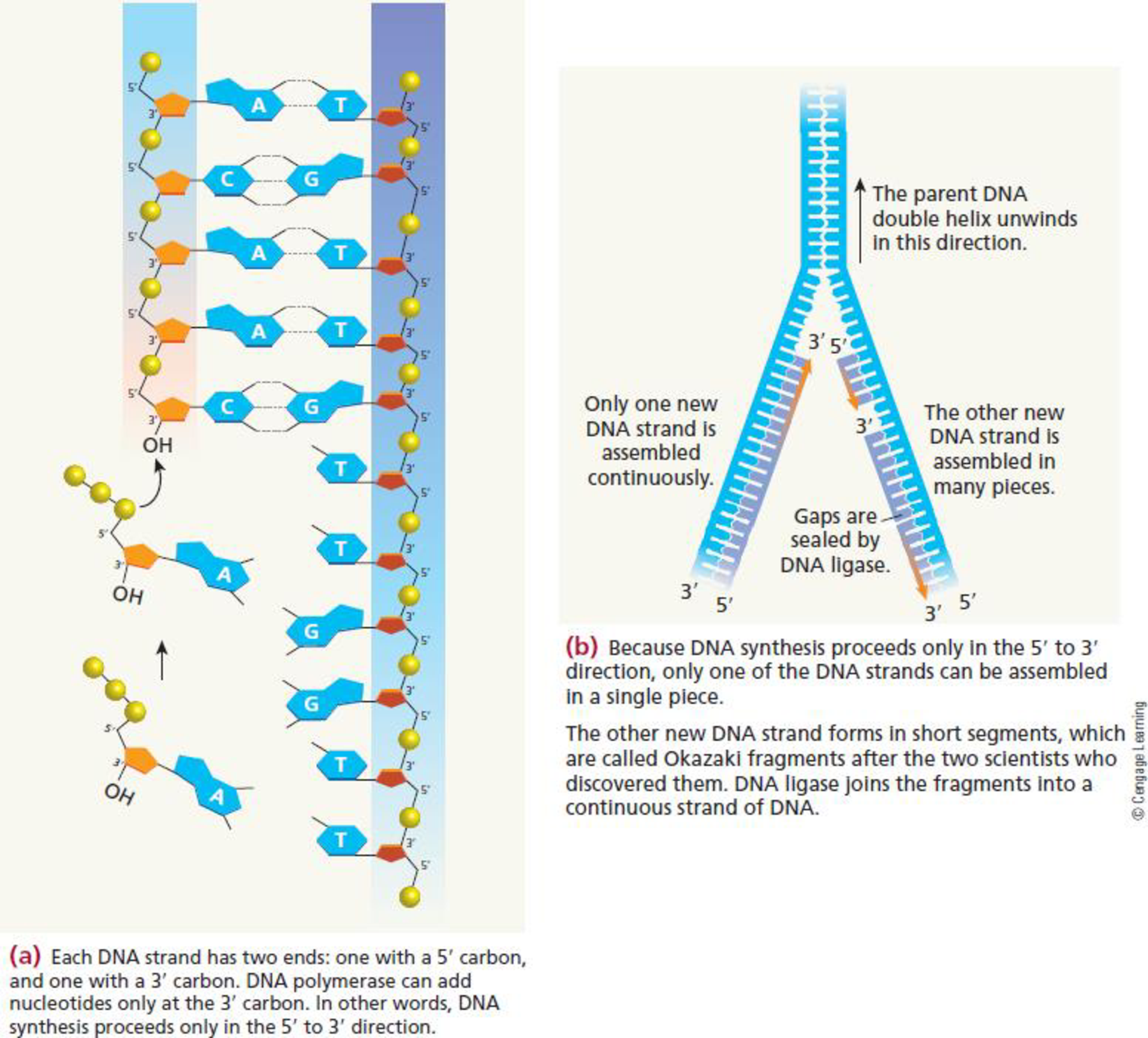
FIGURE 8.13 A close-up look at the process of DNA replication. (a) As the strands uncoil, bases are added to the newly synthesized strand by complementary base pairing with bases in the template strand. The new bases are linked together by DNA polymerase. (b) DNA synthesis can proceed only in the 5′ →3′ direction; newly synthesized DNA on one template strand is made in short segments and linked together by the enzyme DNA ligase.
Trending nowThis is a popular solution!

Chapter 8 Solutions
Human Heredity: Principles and Issues (MindTap Course List)
- What is the difference between the leading strand and the lagging strand in DNA replication? There are different DNA polymerases involved in elongation of the leading strand and the lagging strand. The leading strand is synthesized continuously in the 5' → 3' direction, while the lagging strand is synthesized discontinuously in the 5' → 3' direction. The leading strand requires an RNA primer, whereas the lagging strand does not. The leading strand is synthesized in the 3' → 5' direction in a discontinuous fashion, while the lagging strand is synthesized in the 5' → 3' direction in a continuous fashion.arrow_forwardConsider the following segment of DNA, which is part of a linear chromosome: LEFT 5’.…TGACTGACAGTC….3’ 3’.…ACTGACTGTCAG….5’ RIGHT During DNA replication, this double-strand molecule is separated from the right to the left into two single strands and the replisome is moving from the right to the left of the segment. ___________ should be the template for the lagging strand synthesis. neither of the two strands the bottom strand both top and bottom strands the top strandarrow_forwardTelomerase is a reverse transcriptase enzyme that carries its own RNA molecule (Figure 1(a)). The telomerase enzyme attaches to the end of the telomeric region (Figure 1(b)) for DNA replication. (i) Indicate by drawing where the RNA of Telomerase binds to the telomeric region. W, X, Y, and Z are the ends of the DNA and RNA strands respectively. Identify ends of DNA's X, Y, and Z shown in Figure 1(a) & (b). (ii) (a) Telomerase -AAUCCCAAU- ITTAGGGTTAGGGTTAGGGTTAGGGTTAGGGTTAGGG-W' ПAАTСССААТСССААТСССАА-Х (b) Telomeric DNA Figure 1arrow_forward
- The figure below depicts various elements of the eukaryotic replication machinery in action. Enter the name for the protein depicted by each box. Box A Box B Box C Box D Box E Box F DNA polymerase on lagging strand (just finishing an Okazaki fragment) F Maintains polymerase association with DNA Enzyme extends separation of DNA strands Synthesizes RNA fragments that hybridize to DNA Relaxes supercoiled DNA ahead of replication fork Maintains DNA is single stranded state Promotes binding of processivity factors to DNA Newly synthesized strand pocoar Leading-strand template A New Okazaki fragment RNA primer E Lagging-strand template DNA polymerase on leading strand B C D Saaragon - Next Okazaki fragment will start here Parental DNA helixarrow_forwardWhy is DNA replication is considered a semi-discontinuous process? Explain in detail.arrow_forwardMatch the enzymes involved in DNA replication with their function. Primase [ Choose ] [ Choose] Synthesizes short RNA segment to initiate new DNA strand Helicase Main enzyme that extends RNA primer by adding DNA nucleotides to it Stabilizes single-stranded DNA Relieves over-winding of DNA ahead of the replication fork Removes RNA primers preceding Okazaki fragment and replaces RNA nucleotides with DNA nucleotides Single-stranded binding proteins Unwinds DNA helix Synthesizes the ends of the linear chromosome Seals nicks between adjacent DNA segments DNA polymerase III [ Choose ] DNA polymerasel [ Choose ] DNA Ligase [ Choose ] Topoisomerase [ Choose ]arrow_forward
- Consider the following segment of DNA, which is part of a linear chromosome: LEFT 5’.…TGACTGACAGTC….3’ 3’.…ACTGACTGTCAG….5’ RIGHT During DNA replication, this double-strand molecule is separated from the right to the left into two single strands and the replisome is moving from the right to the left of the segment. The replisome is approaching to a chromosomal end on the left. Considering this left chromosomal end, if without telomerase, the newly synthesized daughter DNA of ___________ will be shortened? neither top or bottom strand both top and bottom strands the top strand the bottom strandarrow_forwardDNA Replication occurs on both prokaryotes and eukaryotes. Although they have a similar genetic flow, there are small differences in between. What are the differences of DNA replication in prokaryotes and eukaryotes? What is/are the major difference/s?arrow_forwardWhat proteins are crucial for creating and maintaining DNA replication forks? Choose the best explanation. Question 2 options: Helicase creates the replication fork; primase keeps the single strands from closing shut. Helicase creates the replication fork; single-strand binding proteins keep the single strands from reuniting. Ligase creates the replication fork; DNA polymerase II keeps the single strands from reuniting. Helicase creates the replication fork; ligase keeps the single strands from closing shut.arrow_forward
- DNA polymerase occasionally incorporates the wrong nucleotide during DNA replication. If left unrepaired, the base-pair mismatch that results will lead to mutation in the next replication. As part of a template strand, the incorporated wrong base will direct the incorporation of a base complementary to itself, so the bases on both strands of the DNA at that position will now be different from what they were before the mismatch event. The MER-minus strain of yeast does not have a functional mismatch excision repair system, but it has normal base excision repair and nucleotide excision repair systems. Which of the following statements is correct about differences in the mutation spectrum between MER-minus and wildtype yeast? More than one answer is correct. Options: More point mutations will arise in MER-minus yeast. Fewer point mutations will arise in MER-minus yeast as compared with wildtype. Of the total point mutations that…arrow_forwardHuman Fbh1 helicase is important in the process of DNA replication. When a mutation occurs during the production of Fbh1, the result is a mutant Fbh1 that binds at the replication fork and prevents any helicase protein from attaching to the strand. Based on this information and the image shown, what would happen during DNA replication if this mutant helicase were present? A - Topoisomerase would unwind the DNA and an RNA primer would attach to the DNA molecule and initiate replication. The process would then stop at the blue triangle because helicase is needed to separate the strands of DNA. B - Topoisomerase would unwind the DNA, but then the process would stop at the blue triangle because helicase, the RNA primer, would not be able to attach to the DNA molecule and initiate replication. C - The process would begin at the blue triangle when topoisomerase unwinds the DNA and an RNA primer attaches to the DNA molecule and initiates replication. DNA polymerase would begin the synthesis…arrow_forwardDNA polymerases are processive, which means that they remain tightly associated with the template strand while moving rapidly and adding nucleotides to the growing daughter stand. Which piece of the replication machinery accounts for this characteristic? Helicase Sliding Clamp Single Stranded Binding Protein Primasearrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
