
Concept explainers
A second strain of dwarf plants has a different mutation of the same gene identified in Problem
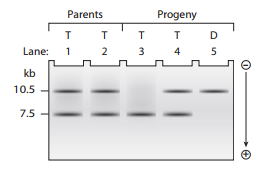
What mutational mechanism is most likely responsible for the production of abnormal DNA fragment length corresponding to this mutant allele? Explain your reasoning.
In comparison to the length of mRNA from the normal allele, will the mRNA from this mutant allele most likely be longer, shorter, or about the same length? Explain your answer.
Want to see the full answer?
Check out a sample textbook solution
Chapter 10 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
- Cyndi grows thousands of B. rapa plants from seed and carefully watches for any unusual plants that might be a new mutation. She has been watching a small seedling whose leaves continue to develop, but the stem is not elongating. The result is a cluster, or rosette, of leaves sitting just above the soil surface. After 15 days the plant is beginning to produce flowers but is still a rosette. Cyndi is encouraged and hypothesizes that rosetteis a recessive mutation. -Describe the first cross she should make and the predicted results. Because B. rapa is an annual plant, the one rosette individual dies. Cyndi would like to continue her study of this mutation, but all od the offspring from her original cross were normal. There are no more plants with the rosette mutation. -What can Cyndi do to continue her investigation of rosette? What would be the results of these "next steps"? -What results would you expect if Cyndi crosses her new generation of rosette plants with a known heterozygous…arrow_forwardThe DNA of every individual in the pedigree shown in image B (below) has been sequenced at the causative locus, all the non-shaded individuals are wild type apart from III.1 and III.6. III.1 and III.6 have both been proven to have the causative allele for the condition but they do not exhibit any of the phenotypic signs or symptoms. Based on this pedigree, what is the level of penetrance for the condition? Please give your answer as a percentage to one decimal place, give the number only, no percentage symbol. Given the information above I calculate the level of penetrance seen in image B to be "Blank" 1 percent.arrow_forwardThe DNA of every individual in the pedigree shown in image B (below) has been sequenced at the causative locus, all the non- shaded individuals are wild type apart from III.1 and III.6. III.1 and III.6 have both been proven to have the causative allele for the condition but they do not exhibit any of the phenotypic signs or symptoms. Based on this pedigree, what is the level of penetrance for the condition? Please give your answer as a percentage to one decimal place, give the number only, no percentage symbol. ANSWER: Given the information above I calculate the level of penetrance seen in image B to be Blank 1 percent. A KEY Homozygous Homozygous Heterozygous Heterozygous Wild Type Male Female Male Female Male Note: Completely red symbol denotes an individual exhibiting the phenotype of interest CI || III IV V 1/4 1/2 1/2 1/2 1/2 Wild Type Female 1/4 1/2 Affected Known carrier Affected female Normal female Affected male Normal male D ●●●arrow_forward
- Consider a maize plant: Genotype C/cm ; Ac/Ac+ where cm is an unstable colorless allele caused by Ds insertion. What phenotypic ratios would be produced and in what proportions when this plant is crossed with a mutant c/c Ac+/Ac+? Assume that the Ac and c loci are unlinked, that the chromosome-breakage frequency is negligible, and the C allele encodes pigment production.arrow_forwardThere are two genetic disorders that result from mutation in imprinted genes: Prader-Willi syndrome and Angelman syndrome. Prader-Willi syndrome results from deletion of region 15q11-q13, which in healthy individuals is a region imprinted such that only the paternal copy is expressed. In the pedigree above, individual I-1 is heterozygous for a deletion of region 15q11-q13 and does not have Prader-Willi syndrome. Individuals I-2 and II-1 are both homozygous wild type for the region. Which individuals in the pedigree might have Prader-Willi syndrome? (Who could potentially have the syndrome, based on what alleles it is possible for them to inherit and express?) Question 9 options: Only II-2 could have Prader-Willi syndrome III-1 could have Prader-Willi syndrome in the presented pedigree; II-2 could only have had it if she were male Both II-2 and III-1 could have Prader-Willi syndrome II-2 could have…arrow_forwardYou are attempting to genotype a series of cells at gene A through restriction enzyme digestion. You know that the wild type allele “A” has 2 restriction enzyme cutting sites, creating bands that are 100bp, 200bp, and 300bp. A mutation creating allele “a” removes one of these cutting sites, creating bands that are 100bp and 500bp. How many separate bands would you expect to see on a gel for a cell that is heterozygous Aa? please show work. thanksarrow_forward
- You are studying four linked genes located on chromosome 2 in the fruit fly Drosophila melanogaster: adp (which contributes to obesity), b (which contributes to body color), pr (which contributes to metabolism), and vg (which contributes to wing formation). A series of crosses between pair-wise combinations of these mutations yielded the following recombination frequencies between the indicated loci: pr and adp 9% adp and b 6% pr and b 3% vg and b 10% vg and pr 7% What is the genetic distance in map units (cM) between the adp and vg loci? 16 9 6 4arrow_forwardCompared to the normal A allele, the disease-causing allele in sickle cell anemia (S allele) is missing an MstII restriction site. On a Southern blot of genomic DNA cut with MstII and hybridized with the probe shown on the diagram below, a person with sickle anemia, carrying two S alleles, will show Choose an answer below: a single band at 1.1 kb. a single band at 1.3 kb. a single band at 0.2 kb. one band at 0.2 and one at 1.3 kb. one band at 1.1 and one at 1.3 kb.arrow_forwardRecombination frequencies between four genetically-linked loci in corn are shown in the following table: Loci Recombination Frequency (%) R and Q 45 W and Q 60 R and W 15 Q and L 10 L and R 35 What is the order of the genes on the chromosome? (note: The same answer can be represented forward or backwards. e.g. A B C D = D C B A) RQWL LQWR QWLR QRLW WRLQarrow_forward
- Two morphotypes of the newly discovered plant in Mt. Banahaw de Lucban were tested for linkage of three genes such as the presence of tendril (t+), dense trichomes (d+), and the presence of secretory cells (sc+). The loci for the mutant genes have been mapped and are separated by the following map distances: t and d = 20CM; d and sc = 12CM. The interference between these genes is 0.4. The first morphotype is located at the lower elevation and is characterized by the absence of tendril, sparse trichome, and absence of secretory cells. On the other hand, the second morphotype is located at the higher elevation and is characterized by the presence of tendril, dense trichome, and presence of secretory cells. Further genetic analysis showed that the morphotypes were true breeding. The two morphotypes were intercrossed and the resulting F1 is testcrossed with the first morphotype. The cross resulted in 1800 progeny. Give the genotypes, phenotypes, and the expected numbers of phenotypes in…arrow_forwardRecombination frequencies between four genetically-linked loci in corn are shown in the following table: Loci Recombination Frequency (%) L and Q 20 Q and R 50 R and L 30 Q and W 13 L and W 7 What is the order of the genes on the chromosome? (note: The same answer can be represented forward or backwards. e.g. A B C D = D C B A) LQWR RQWL LRQW QRLW RLWQarrow_forwardYou have isolated 8 mutants in yeast that fail to grow on minimal media plates but do grow when they are supplemented with Arginine. You know that Arginine is synthesized in a biochemical pathway within wild-type yeast, but you do not know how many gene products it takes for the pathway. You have all of the lines as both a and a cells and mate each strain to each other in pairwise crosses and plate them on minimal media to see if they grow. You obtain the following results with (+) representing growth, and (-) indicating no growth: a 1 5 1 a 4 5 6 7 8 How many genes are represented? O 1 3 7 O Cannot tell from the data a + + + + + • + + i 0 +, + + + • + + 7 + + + + + , . + + + + + m + + + + + + + 2 + + + + + i + -I + + . . + + +arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





